Charged Based Protein Separations
Separations were performed at 2 different salt concentrations with equal volume filtrate and retentate (to avoid dilution problem). Separation time was 40hrs rather than the usual 24.
The lower salt concentration run has a lower cutoff – which is just like what we see in the DNA separations. Remember though that proteins have different charges and shapes unlike DNA and can complicate separations.
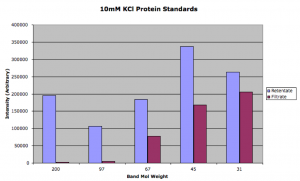
Here is the densitometry data:
You can see that the 100mM was much closer to equilibrium than the 10mM sample. We can use the filtrate/retentate ratio as Paul did in this post to visualize the separation:
While it’s not quite as pretty, especially since we don’t have as many data points, it is possible to observe the saturation and exponential form as shown in the DNA data.
By my rough exponential fit (only to the last three points of the 100mM series), I calculate the 1/e value to be 115 kDa for 100mM KCl and 54 kDa for 10mM KCl.
The pH of the 100mM KCl is reading at about 6.9. This is higher than all of the isoelectric points, which means all of these proteins are negatively charged.
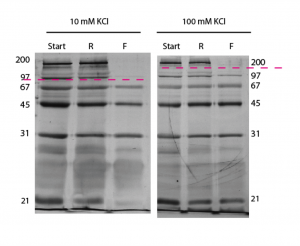


Is the exclusion of the 21 kDa band in the 10mM sample just a staining artifact, or is it a heavily charged molecule? Do you know the amount of excess charge on all these molecules, and are they similar enough to ignore when comparing their diffusion rates through the membrane? I’m not sure if the total charge or the charge/mass ratio is most critical for experiments like this.
This is very nice Jess. Showing the influence of salt on all three species (gold, DNA, protein) elevates the signifcance of this work and makes a rather elegant looking story. I hope you are seeing some of the benefits of our simple pore structure, so that tortuous path polymer membranes do not have this sensitivity to salts. We shall see.
There is no inertia to worry about at the length scale of proteins, and so mass isn’t important per se. It is only important becase size scales with it. I’m imagining that the size and charge work together to determine the effective size. If we knew the isoelectric points of the proteins here it would tell us something about the similarity of charges in this ladder. This is something that the Proteomics Center might be able to determine by isoelectric focusing if we dropped off our ladder with Alan Friedman. I don’t think its a priority right now, but we might like such data down the road.
Chris, that looks more like a staining artifact to me. I only had one experiment for the 10mM work because the second discolored. There will be more repeats up soon so we should be able to tell for sure.
I think the pI should be known for all of these proteins. I’ll look it up and post in above.
How will the magnitude of the excess charge vary with pH? Is it all or nothing as you cross the pI or does charge build as you move away from the pI? The reason I ask is that the protein that shows the greatest change in your experiment is also nearly at the pI. If it follows that this is also the least charged molecule, it makes the charge argument slightly less convincing. If the molecule were neutral, would you expect the salt concentration to affect transport, based on our current theory?
Proteins have many ionizable groups and are zwitterionic at their pI. Over the pI there will be an increase in negative charge as more of the groups become deprotinated. I’m sure it’s not an all or nothing type change.
From my count there are about 118 residues that should be negative (Aspartic acid and Glutamic acid have pKas around 4) and 114 that should be positive (Arginine and Lysine have pKas above 10). The 22 histadines will partially be positively charged (pKa 6.1) and some of the 9 cysteines may be deprotinated (pKa 8). The first carboxylic acid will be deprotinated and the polypeptide ends with a proline (where the side group likes the main chain) and will have no charge. To some extent the local environment can change pKas, so it’s not as simple as adding up residues. This molecule should be negatively charged at neutral pH, but it’s hard to tell exactly how much. We could get purified phos b and run it in the zeta sizer to get a zeta potential.
If it is a completely neutral molecule then I wouldn’t think that salt concentration would have any effect on the transport.
However since we are talking proteins we could be experiencing denaturation at different concentrations. Since all of these proteins are usually disolved in 150mM PBS, I wouldn’t consider my 100mM KCl to be a problem. If anything the 10mM KCl should denature proteins. I have shown before that denatured proteins can weave their way through pores that they wouldn’t be able to get through while in native form, so that would ruin the theory that phos b is being denatured.
I like how it’s not just one protein but the characteristics of the whole separation are changed due to the salt concentration. It’s very similar to the changes seen in the DNA work.
Is it possible to determine what kind of losses you are seeing in these experiments? Now that I’ve seen your set-up, I see that you are really minimizing contact with everything but the chip. Perhaps most of the material is accounted for?
Thanks for all the details!
As I think about this more, I’m not sure the total excess charge is as important as the individual surface charges and where they are located. I assume that if you observe these large molecules from different perspectives, some regions will look more positively charged, while other areas will be more negatively charged. For interactions at the nanoscale, I suspect that this nonuniform charge distribution on the molecule’s surface will affect passage through our fairly uniformly charged pore (aperture). Given this complication, the affect of pKa may get washed out, so a very large pH difference will be required to see a change in transport. I think proteins are fairly complex, so we have to be careful in formulating a theory. On the flip side, no one really has all the answers either, so we should be cut a little slack on a simplified theory.