DLS data on A3G
These experiments were conducted during the week of 5/10/2010 – 5/14/2010.
To recap on this project, Barrett and I have been working with Dr. Harold Smith in the hopes of developing a drug screening test that consists of small-scale diffusions. In these experiments, we are working with APOBEC3G (A3G), a protein-RNA complex that is active in its dimer form but inactive in its natural multimer form. We hope to be able to separate the dimers from the multimers by way of pnc-Si arrays.
The specific goal of these DLS experiments is to demonstrate that we are able to break up the A3G multimer with RNAse and then show transport through pnc-Si membranes.
A3G is an elongated complex, but its HMM form is ~20nm and its LMM form is ~14nm. The tricky part is that A3G is prone to aggregation.
Setups were prepared according to the protocols: “Preparing an RNAse-free setup” and “Working with and digesting HMM”
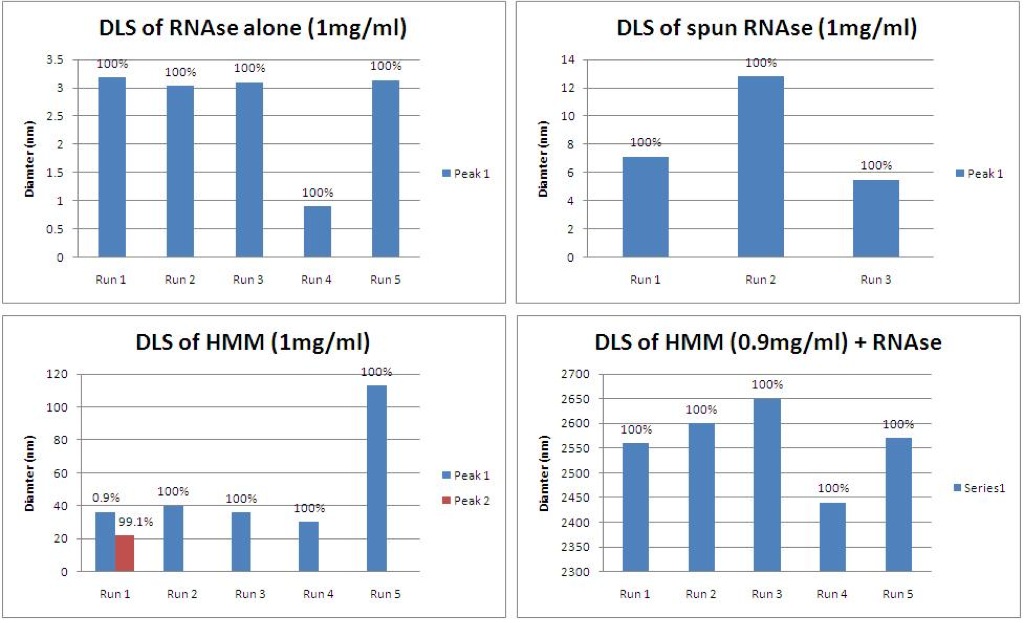
1) The first DLS series investigated HMM (0.9mg/ml) with and without the addition of RNAse. For the experiment where RNAse was added, the setup was allowed to sit on ice for 2 hours to facilitate HMM digestion.
The Malvern conducts several repeat runs but the runs occur successively, inducing a time lapse between readings. The DLS of HMM (on the left) appears to be showing an aggregation of HMM complexes. Runs 1 and 2 show the expected diameter of HMM (~20nm), run 3 seems low to the point that it is around the expected diameter of LMM, and runs 2 and 5 seem dominated by aggregates. The DLS of HMM + RNAse (on the right) appears to be showing a breaking apart of HMM aggregates to the expected ~20nm of a single HMM complex in run 4, followed by a returning to aggregations in run 5.
2) The second series of DLS experiments looked into the RNAse itself, to see if the RNAse was skewing the DLS results. Also, the HMM + RNAse setup was incubated at 37C for 1 hour to investigate the effect of temperature. Barrett and I learned from Dr. Harold Smith that the RNAse was more effective at 37C.
According to the DLS data, the RNAse is much smaller than the HMM complexes, as was expected (top row). Spinning the RNAse seems to have helped remove small particulates that may have been contaminating the RNAse sample. The DLS data for the HMM (bottom left) is still slightly higher than expected, but the data is much more consistent than in the first trial. Run 5 seems to show aggregation of the HMM. Incubating the HMM and RNAse at 37C for 1 hour (bottom right) may have helped activate the RNAse, but it seemed to also greatly increase the aggregation. Or maybe this sample was highly contaminated.
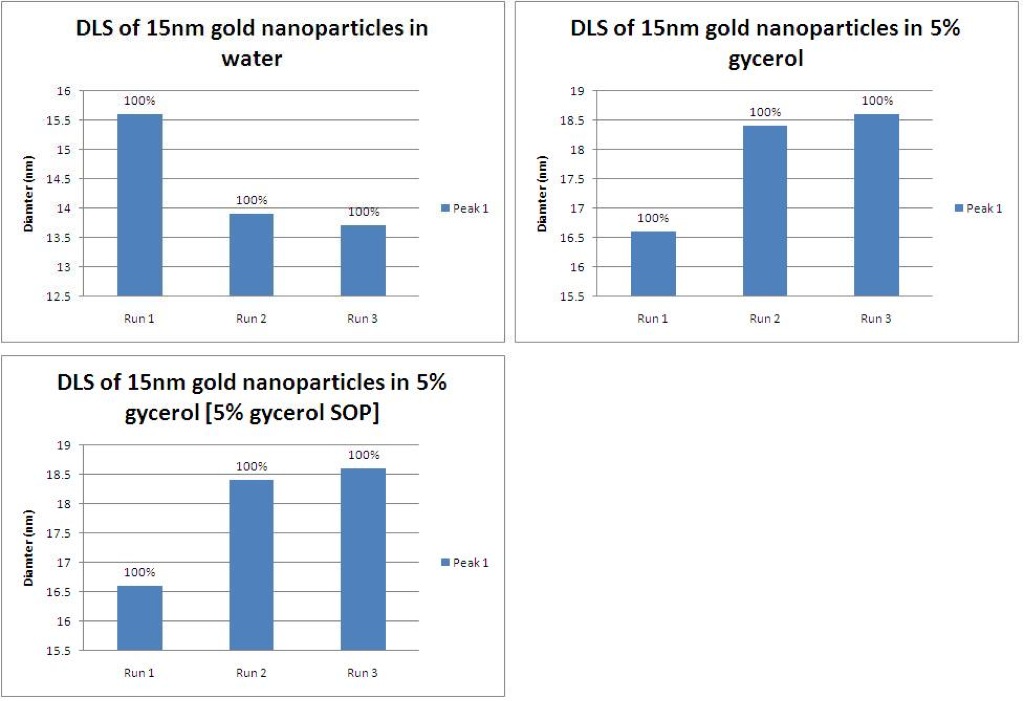
3) Looking at the HMM buffer components, Barrett and I noticed that it is 25% glycerol. Since the viscosity of glycerol is so much higher than that of DI H2O, we decided to test the effect of viscosity on the DLS. Knowing that the gold nanoparticles are 15nm, we can see how the DLS readings change.
The average reported size of the gold nanoparticles in DI H2O is 14.4nm, which is close to the expected 15nm (top left). When 15nm gold particles are placed in a solution of glycerol, the reported size jumps up to almost 18nm (top right). Using these measurements to derive a conversion factor, we made a new operating procedure for 5% glycerol solutions. Using the new operating procedure, the average reported size of the gold nanoparticles was 14.5, which is much closer to the known size and almost identical to the size reading of gold nanoparticles in DI H2O.
4) Using the new 5% glycerol SOP, I again took DLS measurements of HMM with and without RNAse.
These measurements look much closer to the expected values. HMM averaged 24.7nm, before appearing to aggregate quickly thereafter (top left). After 30 minutes, the HMM was a bit smaller than its expected size but it’s possible that the HMM degraded slightly in the warmer room temperature (top right). The RNAse seems to have broken apart the dimer to roughly 8nm (bottom left). Having the setup at room temperature seemed to help activate the RNAse.
In conclusion, the 5% glycerol in the HMM buffer factored into the DLS data, causing the data to be skewed toward higher diameters. Also, the RNAse seems to work best after 30 minutes at room temperature.


It seems like part 1 is irrelevant since the RNAse incubation was not at the optimum temperature. In part 2, we used the suggested RNAse incubation temperature but the data didn’t look great. It doesn’t make sense to me that the size of RNAse alone increased with centrifugation because centrifugation should pull down larger particles, not smaller particles. In part 3, we corrected for the more viscous buffer that we’ve been using – which is important, although it didn’t change the DLS sizing ALL that much.
I’m concerned about the lack of repeatability between different DLS runs. None of the graphs in this post show consistent results between runs. Could HMM be aggregating or disaggregating so quickly that we are picking it up during a DLS run? I wonder if we are introducing contamination somewhere during the experiments, or not cleaning the cuvette adequately for this finicky protein complex.