Fisher Team update: Nanomembranes & lipid particles
In our previous post I relate how our first experience with the membranes encountered some difficulties. These were mostly of two kinds: clogging or braking of the membrane and somewhat arbitrary sizing of the particles. We think we have fixed both.
In our latest experiments we have used manually-extruded particles. These are obtained after the filtration of a polydisperse lipid-micelle preparation through a series of polycarbonate membranes. After passing the prep through a 100nm filter, the zetasizer consistently returns values of 100nm +/- 10nm. This remains the same after the sample has been spun, from 300 to 2000 g. The size is also conserved after diluting the sample, from the original 40mg/ml to 10mg/ml to 2mg/ml. So we are confident with the readings that we obtain when measuring size. That fixes one problem.
The second problem was clogging/breaking membranes with forward centrifugation. We solved this by both diluting the original sample (as I mentioned above) and by finding the sweet spot (speed, g’s) at which to spin down our samples. The 2mg/ml dilution gave us an interesting result, at 4500 rpm. Higher speeds/concentrations result in membrane failure and/or clogging. Slower speeds result in no filtrate.
The result I refer to above is preliminary and pretty simple. We are working on replicates and on performing experiments that involve reverse centrifugations and antibody-particle separation. This is what we got last Friday:
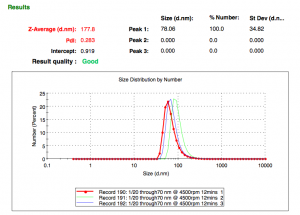
120ul of 100nm particles (2mg/ml) >>> 70-80nm membrane >>> 4500rpm for 12 minutes >>> ~30ul containing ~80nm particles.
We do not know whether we are extruding the lipid particles (forcing ~100nm particles through the nano membrane and changing their size) OR (most likely) selecting the left end of the curve (particles <100nm) and separating the smaller particles from the bigger ones (particles >100nm). Here is the before and after.