Membrane Functionalization: The Demise of Polymer Chemistry and the Rise of KODE
Carrying our membrane functionalization work forward we attempted to work in both the polymer chemistry space and the KODE nonspecific binding space. With the results shown below the KODE molecule pathway has shown much more promise than the polymer chemistry pathway. This, for the moment, has provided the conclusions that fuel the title of this post.
ICAM Experiment
Working with the Waugh lab I made protein G coated chips which we bound ICAM to. I took these chips and attempted to bind a fluorescent secondary antibody (PE Anti ICAM) to them to no avail. Shown in Figure 1 are the images gained from this experiment. The general lack of fluorescence was blamed on potential issues binding protein G to GOPMDS.
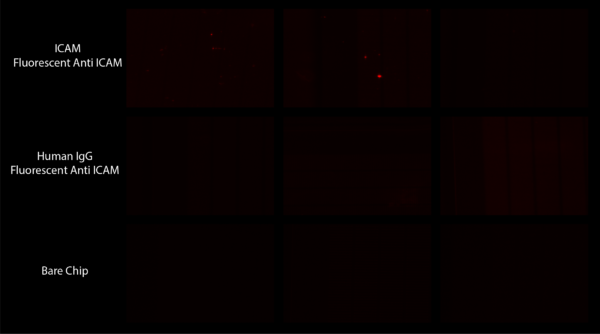

Separately Julie from the Waugh lab attempted to roll cells on these chips to see if they would stick more than on a bare chip. While most of her tests showed no difference, she did notice a pressure difference between one of her ICAM coated chips and a bare chip. She believed this was due to cells adhering strongly within the slits of the chips which she had not seen before.
Fluorescent Human IgG Blocking Experiments
Going back to fluorescent human IgG, we wanted to determine whether or not binding that we saw previously was specific or non-specific and whether or not drying protein G coated chips between cleaning steps affected its binding ability. We did this by setting up an experiment with groups keeping protein G wet, drying protein G, having no protein G, and having no other components other than antibody. The results are shown in Figure 2 and were inconclusive. It was thought that this was do to the fluorescent antibody going bad.

Fluorescent PE Human IgG
To establish whether or not newly order Phycoerythrin (PE) Human IgG was fluorescent and could bind to our chips, we designed an experiment to dip chips into a solution of PE IgG to nonspecifically bind antibody to the surface. The resulting images were black, showing that antibody does not nonspecifically bind to the surface of our chips.
Continuing to address this issue, we designed an experiment after a previously successful one with the addition of showing a group having no nonspecific binding of antibody to our chips. Unfortunately as shown in Figure 3 the results were not the same as they were before, instead showing no fluorescence for any group.
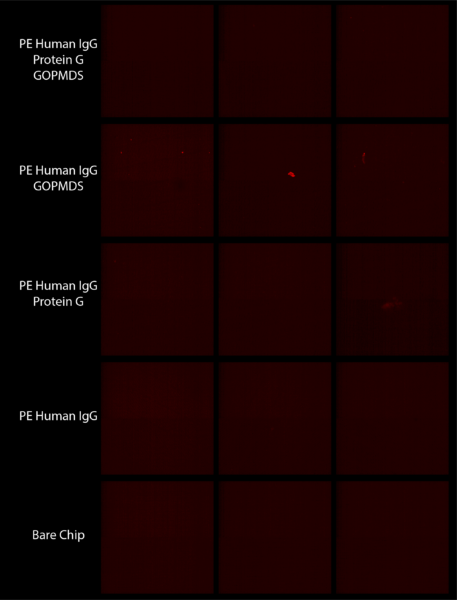

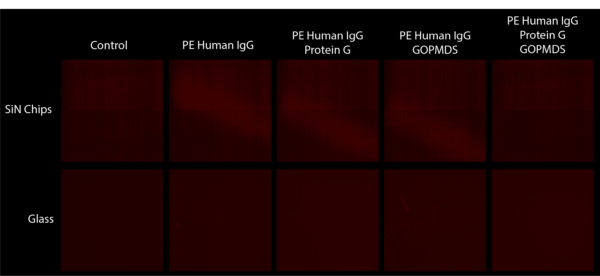
To test our reagents, we completed the same experiment on glass slides and chips. As shown in Figure 4 this again did not work. These results suggest that either one of the reagents is not working properly or the process is fundamentally flawed.
KODE FITC Coating
Attempting a different functionalization path we tried to coat our chips with KODE molecules. To verify coating, we coated our chips with different concentrations of FITC conjugated KODE and imaged with an epifluorescent and confocal scope. As suggested in literature, we imaged the chips without cleaning and kept them wet. These images showed scaling fluorescence intensity as concentration increased and are shown in Figure 5. When these chips were then washed and re-imaged, fluorescence was not seen on any chips. These results told us that KODE molecules may or may not be coating the surface of our chips but regardless they are not very stable.

In a separate experiment I coated silicon nitride and glass chips in a solution of 50 micromolar KODE FITC molecules but allowed the coating to dry onto our membranes. Shown in Figure 6 this led to widespread fluorescence across both silicon nitride and glass chips. Somewhat promising is that the fluorescence remained after a gentle wash of the chips. Figure 7 shows the coating of an entire chip and Figure 8 shows the same chip after a gentle wash.
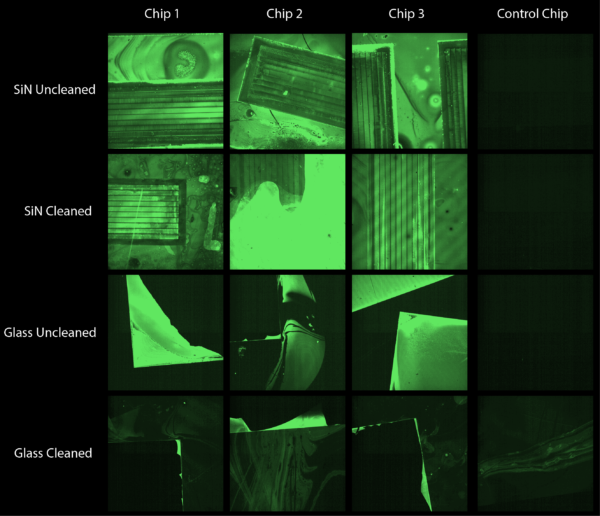
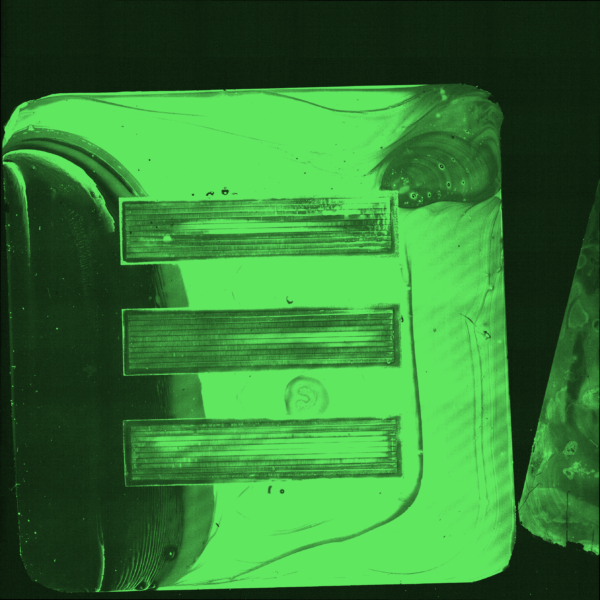



Precision Pico-droplet Printing
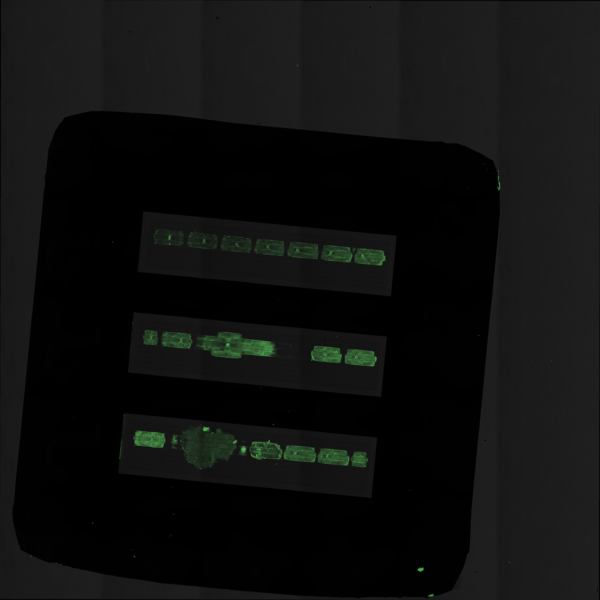
Working with the Miller lab, we proceeded to precision print the same concentration of KODE FITC into the trenches of our 3 slot chips. Shown in the Figures below we can see that we were able to demonstrate what happens in different scenarios of accuracy and volume. Each droplet was roughly 300 picoliters and the same concentration of KODE FITC was used in each experiment (50 microliters).
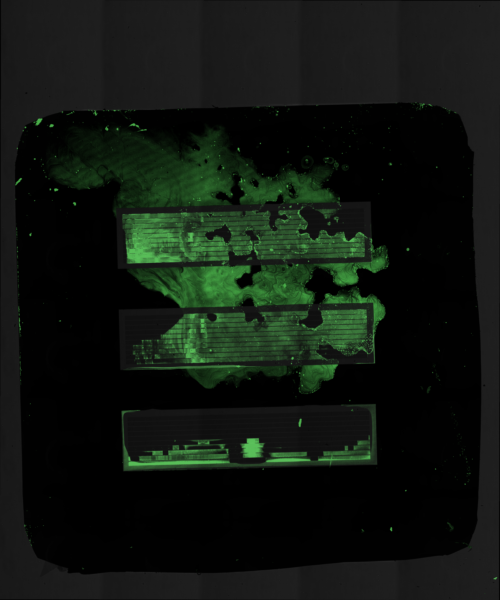



Moving Forward
With these results we are going to move forward with KODE molecule coatings and put polymer chemistry coating on the back burner. I will start by attempting to make the coatings more uniform and determining their stability under various conditions. In addition we will attempt to determine the functionality of these coatings by binding staphylococcus aureus to hyaluronic acid KODE coated chips.
Hi Michael
This is good stuff. What was your clean here? You are at the top end of the concentration recommended for KODE in general and you may be ripping off a multilatered “splat” rather than a good monolayer. This might give you a strong signal that would mean even a residual monolayer after a wash might look like little to no staining. What was your optical set up to take these images? Was the fluorescence visible by eye?
Hey Kevin,
The clean was a gentle wash in DI water. That is a good point about the initial versus cleaned signal. These images were taken on an Andor Dragonfly confocal scope with a 10X objective and stitched together using their fusion software. Fluorescence was not visible by eye.
Michael