hiPSC Differentiation into Brain Pericyte-like Cells, Round 2
Introduction
Breakdown of the blood-brain barrier (BBB) is one of the earliest signs of sepsis and is linked with long-term cognitive impairment. As animal models poorly recapitulate human sepsis, alternative approaches are direly needed to understand sepsis-associated brain injury and develop strategies that protect against cognitive decline. To fill this gap, we are developing a microfluidic model of the human blood-brain barrier (µSiM-hBBB) to elucidate mechanisms of BBB breakdown during sepsis. The first version of the µSiM-hBBB will be developed using human induced pluripotent stem cells (hiPSC) to derive two cellular components of the neurovascular unit: brain microvascular endothelial cells (BMECs) and pericytes. While BMECs are the main component of the BBB, pericytes are key support cell of the neurovascular unit (NVU), along with astrocytes and microglia. Pericytes not only help stabilize the BBB, but are key responders during inflammation and help guide immune cell transmigration.
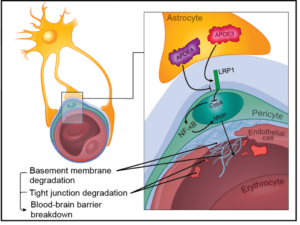
Several protocols have been developed to differentiate pericytes from hiPSCs, however, only recently, a protocol was established to develop brain pericytes following the cells’ developmental pattern (Stebbins et al., Neuroscience 2019). This method drives hiPSCs towards neural crest stem cells (NCSC, p75-NGFR+HNK-1+) and then guides them into NG2+PDGFRβ+ “brain pericyte-like cells”. This post summarizes my second attempt at replicating this protocol in our lab.
Methods
For full details, see hiPSC Differentiation into Neural Crest Stem Cells and Neural Crest Stem Cell Differentiation into Brain Pericyte-like Cells.
D15
- Sort p75-NGFR+ NCSC via magnetic-activated cell sorting (MACS, equipment and reagents from Miltenyi). Analyze by flow cytometry to determine sorting efficiency.
- After sorting, pellet the eluate at 180 x g, 4°C for 5 min and resuspend in neural crest cell cryopreservation medium (6 parts E6-CSFD medium, 3 parts FBS, and 1 part DMSO) to achieve a concentration of 106 cells per mL.
- Transfer 1 mL cell suspension to each cryovial. Transfer cryovials to a controlled rate freezing device and place in a –80°C freezer overnight. The following day, transfer cryovials to liquid nitrogen.
PC Differentiation
- Thaw vial rapidly in 37°C water bath until only a small ice crystal remains.
- Transfer to a 15 mL tube and add 4 mL DMEM/F12 dropwise.
- Centrifuge the cell suspension 5 min at 200 × g, RT.
- Aspirate supernatant and resuspend the neural crest cell pellet in 1 mL E6-CSFD medium. Count.
- Add to E6-CSFD + 10 µM ROCK inhibitor to a density of 5 × 104 cells/mL and add to uncoated 6 well plate.
- Incubate at 37°C, 5% CO2 for 24 hours.
D16
- The following day, switch medium to Essential Medium 6 (E6) + 10% FBS (Peak Serum).
D17 on…
- Feed cells with E6 + 10% FBS daily. Pass when cells reach 90-100% confluency.
- On D22, seed pericytes (or control support cells) with HUVECs for cord formation assay.
- Coat 4 well chamber slides with Geltrex.
- Prep cells (prep each cell using respective protocol for passaging. Resuspend each cell in 1 ml EGM-2 (Lonza)
- HUVECs
- iPSC-derived Pericytes
- Primary Pericytes (HBVP, ScienCell)
- Primary Fibroblasts (derived from 6-month-old wild-type mice hind paws)
- Prepare cell suspensions:
- HUVEC-only control: Add volume of HUVEC suspension containing 5.34×104 cells per chamber
- HUVEC + Support Cell: Add volume of HUVEC suspension containing 5.34×104 cells per chamber, and volume of support cells containing 16.0×104 cells, per chamber (i.e. a 1:3 ratio of HUVECs to support cells).
- Add 1ml/well of appropriate cell suspension to chamber wells and incubate 24 hrs.
- After 24 hours, image using phase microscopy and fix and stain for NG2 and CD31.
- On D25, extract RNA for qPCR or RNAseq from 5 wells of 6 well plates (Qiagen, RNeasy Plus Minikit). Also extract RNA from undifferentiated hiPSCs (IMR90)-4 as control. Save RNA from 2 wells for RNAseq, reverse transcribe the other three wells using Superscript III Reverse Transcriptase (Thermo Fisher). Using 1 µg of cDNA, run qPCR on QuantStudio3 System with Comparative CT (CT), standard run mode.
Analysis
- Cord Formation Assay
- Phase Microscopy: Analyze images with WimTube, calculate average tube length and tubes/field relative to HUVEC alone.
- ICC: Fix cells and stain for endothelial cell marker, CD31, and pericyte marker, NG2.
- qPCR: Compare mRNA expression between hiPSCs and D25 iPSC-derived brain pericyte-like cells by showing expression relative to GAPDH and normalizing to undifferentiated hiSPCs.
Results
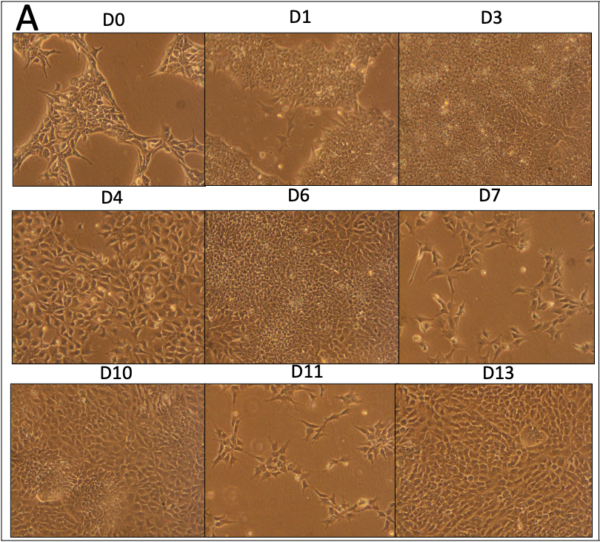
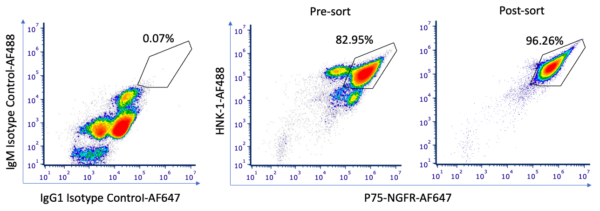
Our NCSC differentiation morphology and MACs data are similar to those reported in Stebbins et al. Importantly, we saw clusters of cells that were p75-NGFR–, which were subsequently sorted out via MACS. While the presort pattern is slightly different than the last round of differentiation, there is a good percent of double positive (P75+/HNK1– population) post-sort. 16 vials of NCSCs were frozen post-sort, which we can use for our next round of differentiation/troubleshooting.

Following MACS, the cells were guided to pericytes, again with morphology Dr. Shusta’s groups. While the growth rate of these cells differed slightly from the last round, the final cells matched well in morphology.
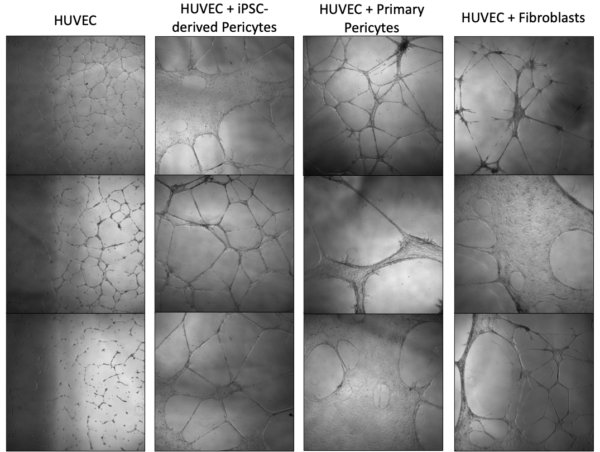
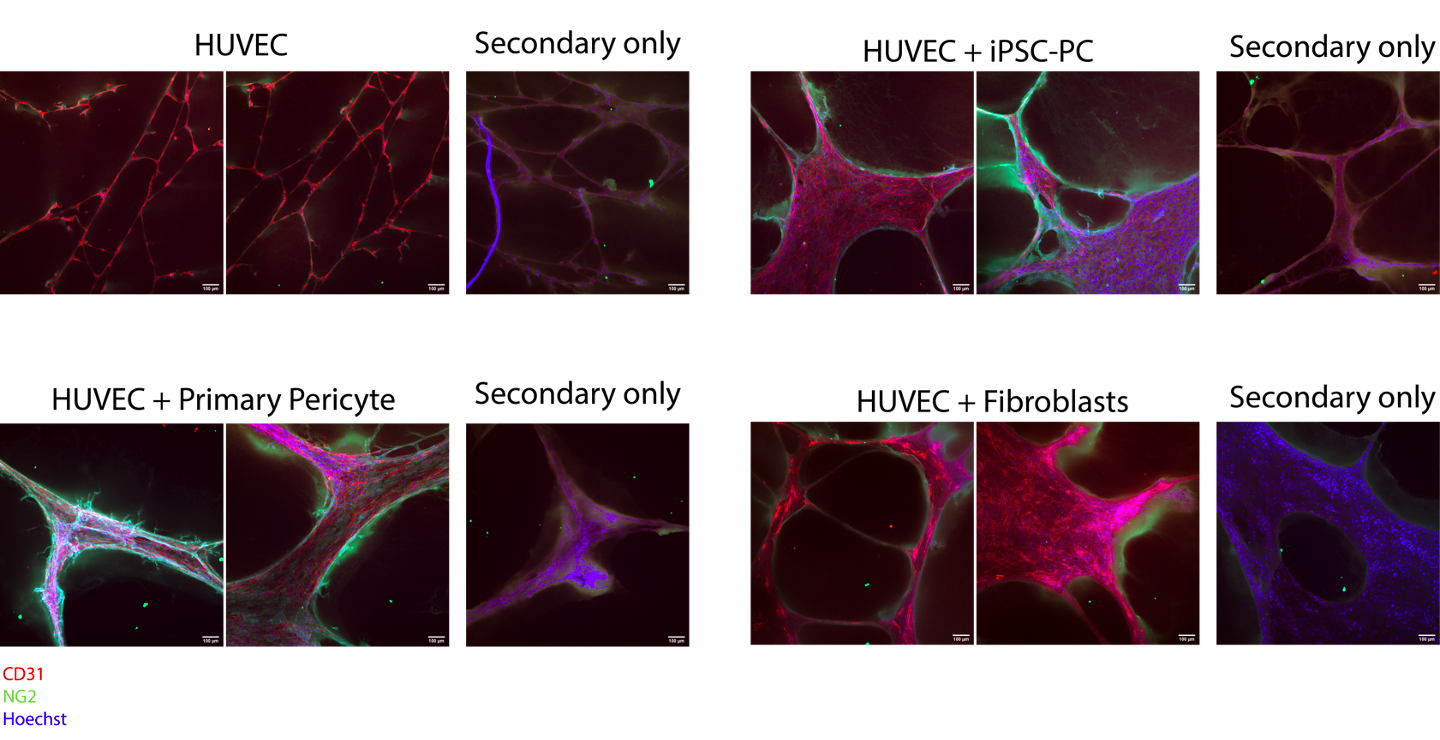
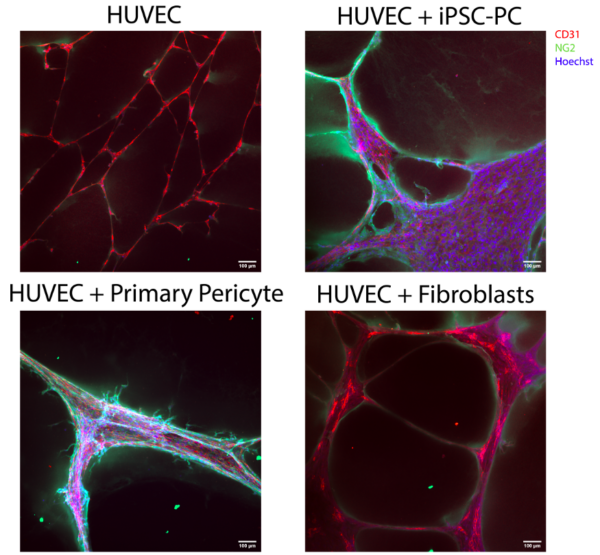
At D22, iPSC-derived pericytes or control support cells were cultured with HUVECs (or HUVECs alone) in chamber slides for a cord formation assay. Note that the positive control may not be fully reliable. I passed these cells earlier and kept in conical in incubator while setting up assay. They began to stick and I had to spin and dissociate with TrypLE. This may have affected their behavior in the assay. However, they still associated very nicely with the HUVECs based on ICC images and appear to stimulate tube formation based on phase images. Comparatively, if not better, the iPSC-derived pericytes stimulate cord formation and associate directly with the cords formed by HUVECs. It is also important to note there was significant green fluorescence background signal, although the intensity is lower than in pericytes associated with HUVEC tubes.

| Gene | Description/Comments | |
| NANOG | Transcription factor that suppresses cell determination factors. Should see no expression in iPSC-PC but not the case | |
| NGFR | “Nerve growth factor receptor” found in NCSC at development, aids in neuronal cell survival and differentiation. Hard to interpret without NCSC (Shusta also had one D25 pericyte-like cell sample that had some retained NGFR expression) | |
| ZIC1 | Possible brain-specific mural cell marker, should be expressed in NCSC, weak expression in iSPC-PC. Hard to interpret without NCSC, but shown to have weaker expression in IMR90-4 | |
| ETS1 | Transcription factor and cranial neural crest marker, role in delamination of NC cells. Hard to interpret without NCSC | |
| PDGFRß | Vascular development, expressed by PCs throughout body and binds PDGF released by ECs (classic pericyte marker) | |
| CSPG4 (NG2) | Chondroitin sulfate proteoglycan and mural cell marker, important in blood vessel formation (classic pericyte marker) | |
| TBX18 | Transcription factor and mural cell marker, expressed throughout body | |
| FOXF2a | Possible brain-specific mural cell marker. Good to see increased expression here | |
| RGS5 | Regulator of G protein signaling-5”, pericyte marker. Variable expression in Shusta data but always similar to hiPSC | |
| ABCC9 | ABC transporter, may distinguish brain pericytes from smooth muscle cells. Shown to have weaker expression in IMR90-4 | |
| KCNJ8 | G protein-activated inward rectifier potassium channel 3, may distinguish brain pericytes from smooth muscle cells. Shusta saw low level of absolute expression in RNA-seq data (ok ours is low) | |
| ACTA2 (SMAa) | Capillary PCs are NG2+SMAa–, contractile (transcript may be expressed temporarily in our cells). Lots of debate on this marker for pericytes, results ok compared to Shusta’s |
Most markers followed the trends seen by Dr. Shusta’s group. However, we saw greater expression of pluripotent marker, NANOG, than expected. We anticipated no NANOG expression in our D25 cells.
Conclusions
While we have mostly positive results for our second round of differentiation, we have some optimizing/troubleshooting to do based on our qPCR data. Our NCSC differentiation morphology and MACs data are similar to those reported in Stebbins et al. Importantly, we saw clusters of cells that were p75-NGFR–, which were subsequently sorted out via MACS. While the presort pattern is slightly different than the last round of differentiation, there is a good percent of double positive (P75+/HNK1– population) post-sort. 16 vials of NCSCs were frozen post-sort, which we can use for our next round of differentiation/troubleshooting. While the growth rate of these cells differed slightly from the last round, the final cells matched in morphology and showed similar gene expressions.
Following MACS, the cells were guided to pericytes, again with morphology Dr. Shusta’s groups. At D22, iPSC-derived pericytes or control support cells were cultured with HUVECs (or HUVECs alone) in chamber slides for a cord formation assay. While I still need to quantify phase images, it is clear both pericyte groups stimulated tube formation of endothelial cells, with greater tube length and reduced number of tubes per field. Further, ICC imaging confirms that both pericyte groups associate directly with the cords formed by HUVECs, although there are differences depending on field of view.
At D25, RNA was extracted from pericytes and undifferentiated IMR90-4 control wells, which were used for qPCR or will be used for RNAseq. While most markers followed the trends seen by Dr. Shusta’s group, we saw greater expression of pluripotent marker, NANOG, than expected. We anticipated no NANOG expression in our D25 cells. Our next step will be to collect RNA from NCSCs to see if they have high expression of NANOG as well. We also need to compare NCSC marker expression in NCSCs to D25-PCs. For our next differentiation, we will use ICC on NCSCs and D25-PCs to see if we have some expression of NANOG in all cells or high expression in a small population of cells. We may need to improve our cells sorting technique or adjust media components to improve differentiation.
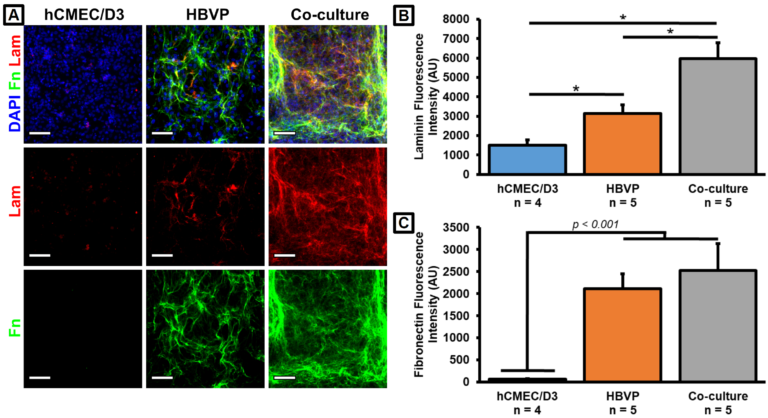
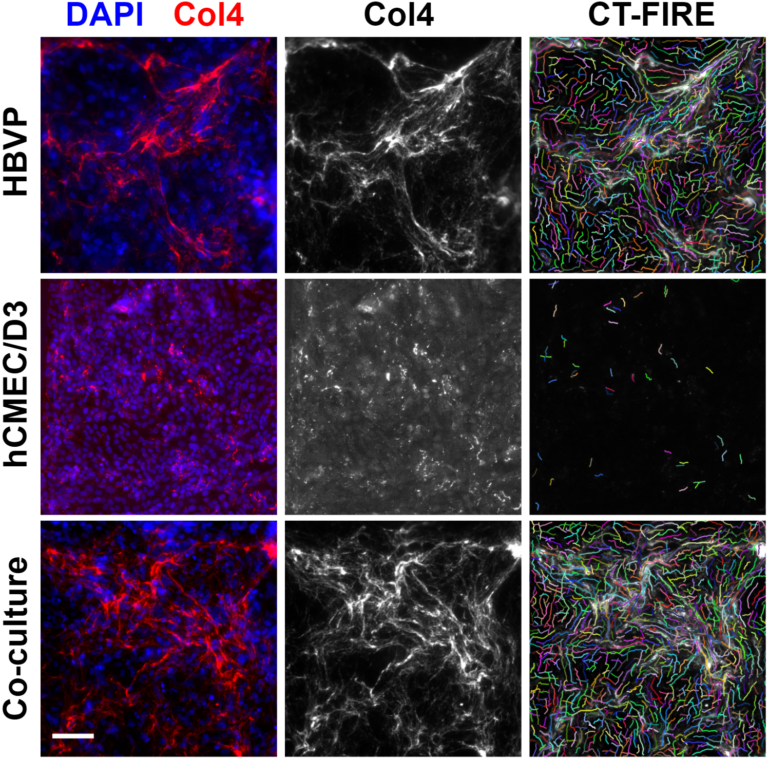
Can you comment on the consequences of elevated pericyte NANOG in our co-cultures? Will it really matter? Will the iPSC experts reject us?
NANOG often indicates the cells have the ability to differentiate into other cell types. Pericytes have some capability to do this already and upregulate NANOG in these situations (see
https://stemcellsjournals.onlinelibrary.wiley.com/doi/full/10.1002/stem.1977), but I would be concerned if they have baseline expression. I don’t want to induce differentiation into a different cell type under conditions they normally would not do so. We would no longer be learning about pericyte contribution to septic responses. Jon might have some more thoughts on this as well