A closer look at separations
UPDATED 12/12/07 – Added data on albumin. In these gels, the band from beta-galactosidase appears to overlap with phosphorylase b making it difficult to determine which protein passes and which is retained.
I have used Mike Bindschadler’s pore imaging software to find the pore sizes of the three membranes I have use to perform succesful separations. The results of the pore processing are compared to the gels in the following images:
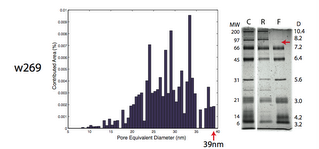
w269 – Pore cutoff around 39nm, separation cutoff around 90kD
w213 – Pore cutoff around 23nm, separation cutoff around 90kD (possibly some much larger pores exist on this membrane)

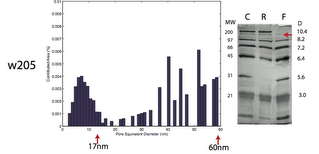
w205 – Cutoff of smaller pore distribution around 17nm but larger pores out to 60nm, separation cutoff higher than 100kD
A note on the three largest proteins in the mixture:
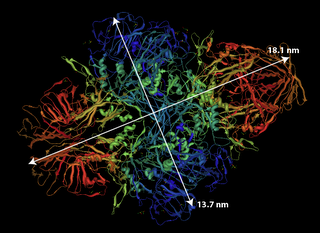
1. Muscle Myosin – Made up of 2 heavy chains each of 200kD each and 4 light chains of 20kD each. Elongated tail measures about 160nm. No pdb file of full molecule. Purification information can be found in Woods et al. J. Biol. Chem. 1963.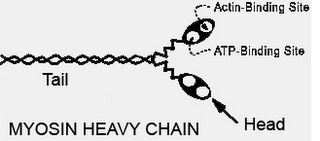 2. beta-Galactosidase – Naturally tetramer a of 464kD. Using the pdb file, dimensions are 18.1 nm by 13.7 nm. In these gels, the band from beta-galactosidase appears to overlap with phosphorylase b making it difficult to determine which protein passes and which is retained.
2. beta-Galactosidase – Naturally tetramer a of 464kD. Using the pdb file, dimensions are 18.1 nm by 13.7 nm. In these gels, the band from beta-galactosidase appears to overlap with phosphorylase b making it difficult to determine which protein passes and which is retained.
3. Phosphorylase b – Monomer is 97.5kD, biologically a dimer. Using the pdb file, dimensions of the monomer are 8.2 nm by 7.3 nm; the dimer would be about 14 nm by 8.2 nm.
4. Albumin – 67kD, dimensions are 8x8x6nm. This protein passed through all three membranes.