Diffusion Separations using w319
My previous post outlined the results of a EQ format separations using membranes from wafer 319. I have followed up that experiment using the same membranes, but in a diffusion based separation. I separated 2uL of 9.3 mg/mL clarified brain extract using a reservoir of 60uL. I allowed the diffusion process to occur overnight, and I collected the samples and ran on a gel. The retentate was diluted to be of similar concentration to the filtrate (about a 30x dilution).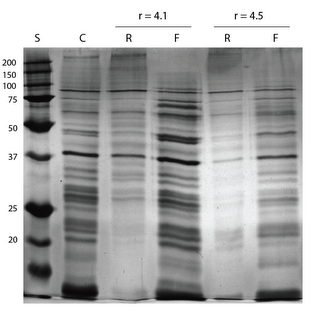
This gel shows a slightly higher cutoff than the previous EQ format experiment. The upper dark band in the brain extract actually appears in the filtrate. The cutoff, while higher, is still very sharp. It is very apparent that there are very few proteins of low molecular weight remaining in the retentate after 24 hours (we do expect this experiment to approach equilibrium, but the volume of the retentate is lower thus giving us a lower total amount of small proteins in the retentate at equilibrium).
The reason for the higher cutoff is unclear. Initially I predicted that the cutoff would be lower, since the proteins are given a “drift velocity” in the flow experiments. I thought the directed motion would help large proteins to approach and pass and a random walk may not be as successful. Since this is clearly not the case, one alternative hypothesis from Jim is that we may be altering the pores more in the flow mode. There may be a higher incidence of large proteins adsorbing to the surface and occluding small pores than in the passive diffusion mode.
To test this hypothesis, we would have to observe the degree and quality of adsorption. We have previously incubated samples in protein and observed using TEM. Could we do this with a sample from a flow based separation? Currently the SepCon membranes will not fit in the TEM goniometer. Is SEM a reasonable substition, or can we not achieve high enough resolution?
The resolution of our current SEM is probably too low, but you are welcome to try. The new SEM should be able to do a better job. There is no way to place the larger inserts in the TEM.
However, I can’t see how the binding could be greater in flow experiments, since you have a much higher protein concentration in the diffusion setup. Seems like this much stagnant protein on a membrane for this long would fully saturate the adsorption sites.
Isn’t this just a consequence of the random walk? Since we are diffusion limited, transport is relatively independent of pore density, so all pore sizes are in play, and the cutoff should match the largest pores. In a flow system, you are limited by the total number of pores at each size, so a small number of large pores would not contribute very much, hence the apparent cutoff would be lower. Of course, our flow system should be a mixture of random walk and flow, so who knows…
The explanation for the higher cut-off is not obvious to me. If we don’t think we can get a good look at the pores, we shouldn’t waste time trying. I’ll dig a bit into sieving theory for diffusion and convection and see if we can rationalize this.
In the meantime, I think the sharpness of the cut-off can be compared to PES membranes, hopefully favorably. To make the story most complete we should do PES in diffusion as well as convection.
It might also be time to break out those gold beads we bought. Seems to me that if the cut-offs and permeabilities are defined by our pore histograms with gold beads, then we could safely point to protein modification of the membranes as the determinant of the new ‘cut-off.’
To document protein binding we should probably be satisfied if we can solidifying the preliminary results you’ve shown before. Again a comparison to PES would be helpful – we can brainstorm an approach that doesn’t involve fluorescence.
So …
1) comparison to PES separations by diffusion and convection
2) analysis of pnc-Si fractionation of gold beads by these two modes
3) protein binding to membrane surfaces
4) Publish!
I like the way Chris put it in his last paragraph. I too had been thinking along these lines for an alternative hypothesis.
Jim/Jess – If big proteins are in fact narrowing the pores only during the convection step, could you take a post-run SepCon and do a diffusion experiment? You are expecting the cutoff to remain low, right? Since it would be hard to get significant protein diffusion in the SepCon format, maybe this would be the time to test the gold bead diffusion.
Sample Protocol:
-Assemble 4 SepCons.
-Let 2 diffuse in the EQ format for 24 hrs (no force)
-Run 2 for 2 hours (hrs 22-24)in the std EQ format
-Remove protein from all four samples and replace retentate solution with gold beads that you expect to pass in one case but not the other (I appreciate this will be the hardest step by far)
i have been wondering, is it possible that the larger molecules are preventing the smaller ones to pass through the pores during convection. this could happen if the larger ones get stuck to the pores and decrease the porosity for the smaller ones. this seems to not happen in case of diffusion as the larger molecules don’t get sucked towards the pores like in convection. this might give rise to the difference in separation of molecules in diffusion and convection methods.