Glial cells can grow as multilayers upside-down
In my 1st post detailing glial cells on pnc-Si transwells, I saw lots of cell clumps above the membrane window (see here). Since these cells were grown in the wells, I thought they could be piling on top of each other in the wells and getting trapped. Here, I decided to flip the sample so that the glial cells grew in the wells, but upside down to see if this cell clumping still occurred. I suspected that cells would fall to the bottom of the 24-well plate instead of forming monolayers.
Here, NG108 cells (P13) were seeded at 50,000 cells/cm2 on the well-side of pnc-Si and then inverted so that they were upside-down in the 24-well plate for growth. TEER was measured and then the cells were stained with Live/Dead on day 7.
PET transwells showed a largely viable, nearly confluent cell layer with some multi-layer clumps. Even though these cells are hanging upside-down, their intercellular adhesion is strong enough to maintain multiple layers away from the membrane surface.
TEER:
TEER of control samples (PET) didn’t change. The pnc-Si results were confusing. Most of the samples stayed right around their day 0 values (that is, there was no increase in barrier function). However, a couple of samples showed ~50% increase in TEER over 1 week. Since there was a fairly confluent monolayer of cells on PET (see above) that didn’t cause an increase in TEER, I didn’t know what to make of this. So I looked at the images:
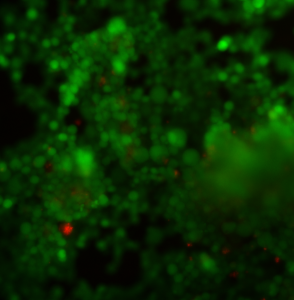
Sample 1 (same sample as in the graph) with the phase contrast channel shown in blue. There is clearly a huge clump of cells away from the membrane, but the cells don’t cover the entire free-standing membrane area.
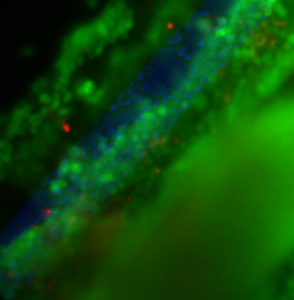
Sample 2 (same as in graph) is below. There are so many cells that the entire image is blurry. It appears that there is a layer several cells deep that covers the free-standing pnc-Si area.
 These drastically different morphologies likely explain the difference in TEER values for these 2 samples. The other samples (4, 5 and 7) exhibited similar coverage differences.
These drastically different morphologies likely explain the difference in TEER values for these 2 samples. The other samples (4, 5 and 7) exhibited similar coverage differences.
How are these cells clumping together when they are growing upside down? Here’s an image to show my working model:
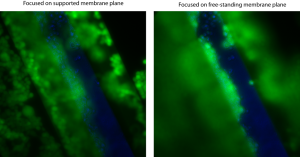 On the left is an image with the focal plane at the supported membrane and on the left is an image with the focal plane at the free-standing membrane. On the left image, cells on the supported membrane and the interface between the well walls and the supported pnc-Si area are both in focus. Clearly, there are cells growing on the sloped well walls. It looks like cells growing on supported pnc-Si form an almost continuous layer with cells growing on the well walls and then down onto the free-standing membrane. Now, this image isn’t perfect since there are very few cells on the upper-right area of supported pnc-Si. BUT, if the cells on either side of the wells (on the supported pnc-Si) form a somewhat continuous layer with cells on the well walls and the free-standing membrane, then dividing cells can stay attached to this cell layer within the wells – thus forming clumps.
On the left is an image with the focal plane at the supported membrane and on the left is an image with the focal plane at the free-standing membrane. On the left image, cells on the supported membrane and the interface between the well walls and the supported pnc-Si area are both in focus. Clearly, there are cells growing on the sloped well walls. It looks like cells growing on supported pnc-Si form an almost continuous layer with cells growing on the well walls and then down onto the free-standing membrane. Now, this image isn’t perfect since there are very few cells on the upper-right area of supported pnc-Si. BUT, if the cells on either side of the wells (on the supported pnc-Si) form a somewhat continuous layer with cells on the well walls and the free-standing membrane, then dividing cells can stay attached to this cell layer within the wells – thus forming clumps.
This phenomenon would certainly be time- and ‘initial cell seeding density’ – dependent. I’m not sure why there are differences in clumping in this experiment. It could be differences in handling since these cells are not very adherent. I need to minimize this clustering in order to minimize the TEER of the glial cells.