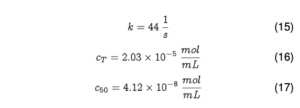ECMO Model Design
While experimental results are still forthcoming, I thought I’d make a post about the nature of the model I’ve constructed for ECMO. This model is the one that my experiments are seeking to validate; so to be clear, this model is founded entirely in theory for now.
ECMO shares some obvious similarities with hemodialysis — blood in an extracorporeal circuit enters a membrane space in which diffusive transport across the membrane is the primary therapeutic mechanism — but from the perspective of a modeler poses a new problem: reaction. Whereas a complete description of hemodialysis can be obtained by modeling transport through convection and diffusion alone, the addition of a reaction term to the governing differential equations complicates efforts to arrive at a concise, closed-form solution.
Fortunately, our collaboration with Professor Al Clark, Jr. equips us with an excellent starting point. In 1985 Prof. Clark first-authored a paper titled ‘Oxygen Delivery from Red Cells’ in which he and his coauthors propose a set of differential equations and corresponding parametric values for describing the unloading of oxygen from red cells into tissue. I have secretly begun to suspect that Prof. Clark is actually a wizard, but that’s for another post.
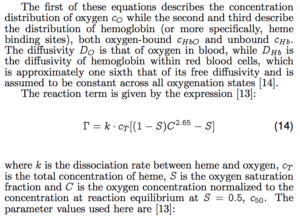
Here, I’ll defer to my recent ICNMM paper to provide the equations and a technical description:
Long story short, we have three simultaneous partial differential equations for oxygen, deoxyheme, and oxyheme, each of which models diffusion and convection as well as an independent reaction term. Someone much smarter than I might be able to solve these equations analytically, but for this problem I go straight to COMSOL.
The Transport of Diluted Species physics module contains a handy reaction term, so my first attempt at this model looked exactly like my hemodialysis model with reaction tacked on: solve for fluid flow, pass that uni-directionally to diffusion (and now reaction) and solve for steady-state solution.
Unfortunately, I learned my first lesson about ECMO the hard way at this point: blood has a ferocious appetite for oxygen. To put it in perspective, at 100% saturation one mL of blood contains approximately 0.2 mL of oxygen at atmospheric pressure. That means that in order to satisfy one mL of blood, about one mL of atmospheric gas must be depleted of oxygen. And going from zero oxygenation to complete oxygenation under atmosphere occurs in less than a minute. All this is to say that reaction in this system occurs very rapidly, and rapid changes mean steep gradients, which don’t play well with finite element modeling!
In order to make this work, then, three things are required: a very dense mesh, a transient simulation with good initial conditions, and a very powerful machine to run it on! These are very resource-intensive simulations, but with enough tweaking and patience, it can work.
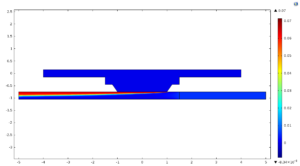
There are two distinct computational domains in this model. The top domain represents the gas phase and contains only diffusive transport of oxygen to conserve computational resources. Because in my experiment there’s no gas flow, just a gas phase of constant pressure, there’s no outlet in this domain. This makes flow very difficult to simulate, so I just cheated and gave the oxygen gas a very high diffusivity in this space so that it’s effectively always at the appropriate concentration everywhere.
The bottom domain is the liquid phase and does contain a fluid flow simulation, which as I mentioned, is solved first and then passed uni-directionally to the Transport of Diluted Species physics. This domain is also the one that contains the reaction term. In my model I’ve separated it into two parts, one before the flow reaches the membrane and one at, around, and after the membrane, again to conserve computational resources since the absence of oxygen in the before-membrane area means there’s no reaction occurring and thus no need for a dense mesh.
The two domains are connected as an “identity pair” rather than a union (union being the default option for assembling domains in COMSOL) and gas transport across them occurs through a thin diffusion barrier boundary pair node which is permissive only to oxygen and models the transport characteristics of the membrane just as they were modeled in hemodialysis.
Of note is that a previous simulation of a similar nature did not include this model of the membrane — instead, I assumed transport across the membrane would occur so readily that the real limiting factor would be the solubility of oxygen in blood. This new model more-or-less confirms that, but only for fairly high gas phase pressures. More work remains to be done to fully characterize under what conditions this assumption becomes reasonable.
One very salient result from this model is the influence of membrane thickness on gas transport. There’s a pretty sexy takeaway under some conditions…
There’s a clear asymptote as the membrane thickness approaches zero where the membrane no longer inhibits transport at all (only diffusion of oxygen in water is limiting,) and nanomembranes approach this point fairly closely. The “asymptote” on the bottom is not a true asymptote, as the flux should continue to decline as you increase membrane thickness indefinitely — the effect is just diminished because it goes as the square of the thickness.
While this general shape is constant regardless of the particular experimental parameters, the specific data on this plot are not strictly valid for all possible conditions. Changes to blood flow rate, gas phase oxygen pressure, hemoglobin concentration, and so on are likely to shift things around somewhat, so more work remains to be done in order to create a truly robust understanding of the conditions under which nanomembranes behave so favorably.
More satisfying conclusions about ECMO still forthcoming.
I’ve uploaded the model .mph file here so that it’s available should anyone want to revisit it at any time. It was designed and executed on COMSOL Multiphysics 5.0, but may be compatible with other versions. Unfortunately, WordPress doesn’t allow .mph files for “security reasons” so I had to give it a different file extension. Just manually change the extension back to “.mph” after downloading and it should work fine.