Nanovesicle Rigidity: Can Exosomes be Modeled as Rigid Particles?
[latexpage]
This an extremely important topic that has come up multiple times in discussions within NRG as well as during my qualifying exam. The question is: are gold nanoparticles a good model for exosomes? The reason that this question comes up is because exosomes are biological material and as such they have some vastly different properties than gold nanoparticles, which are inorganic. The most important property that has been brought up for consideration, however, is the property of deformability. In doing all of my modeling and designing a model system, the question has always been: can exosomes be properly modeled by a rigid particle? This is the most basic assumption, the one that allows to simplify our analyses, making life a lot easier, decreasing the number of frustration-induced, alcohol-fueled benders that I could likely expect in my future if my model system does not correlate well with my experimental system. Therefore, in this post, I set out to determine two things: what is the deformation of exosomes in our systems and can they be modeled as rigid particles?
To start this adventure, I read a paper that Rick gave me after my qualifying exam. This paper is titled “Competition between Bending and Internal Pressure Governs the Mechanics of Fluid Nanovesicles” and is super convenient with its analysis of the mechanics of nanometer-sized vesicle membranes (Supporting Information). This is also my suggestion for a journal club review, so keep a lookout for that if you are interested. In this paper, the authors give values for the stiffness of nanovesicles and these values can be used to approximate the deformation that a nanovesicle will experience. For 100 nm lipid vesicles, they found that the stiffness is ~0.021 N/m, which, if you aren’t familiar with the scale of these things (like I wasn’t), is actually quite stiff. Red blood cells have stiffness values on the order of 7-15 μN/m or 3 orders of magnitude less. This stiffness can then be used to approximate a deformation simply using the force that is being applied to these particles.
In order to calculate the force on the particles, I did a simple approximation from pressure using the following equation:
\begin{equation}
F = \Delta P * A
\end{equation}
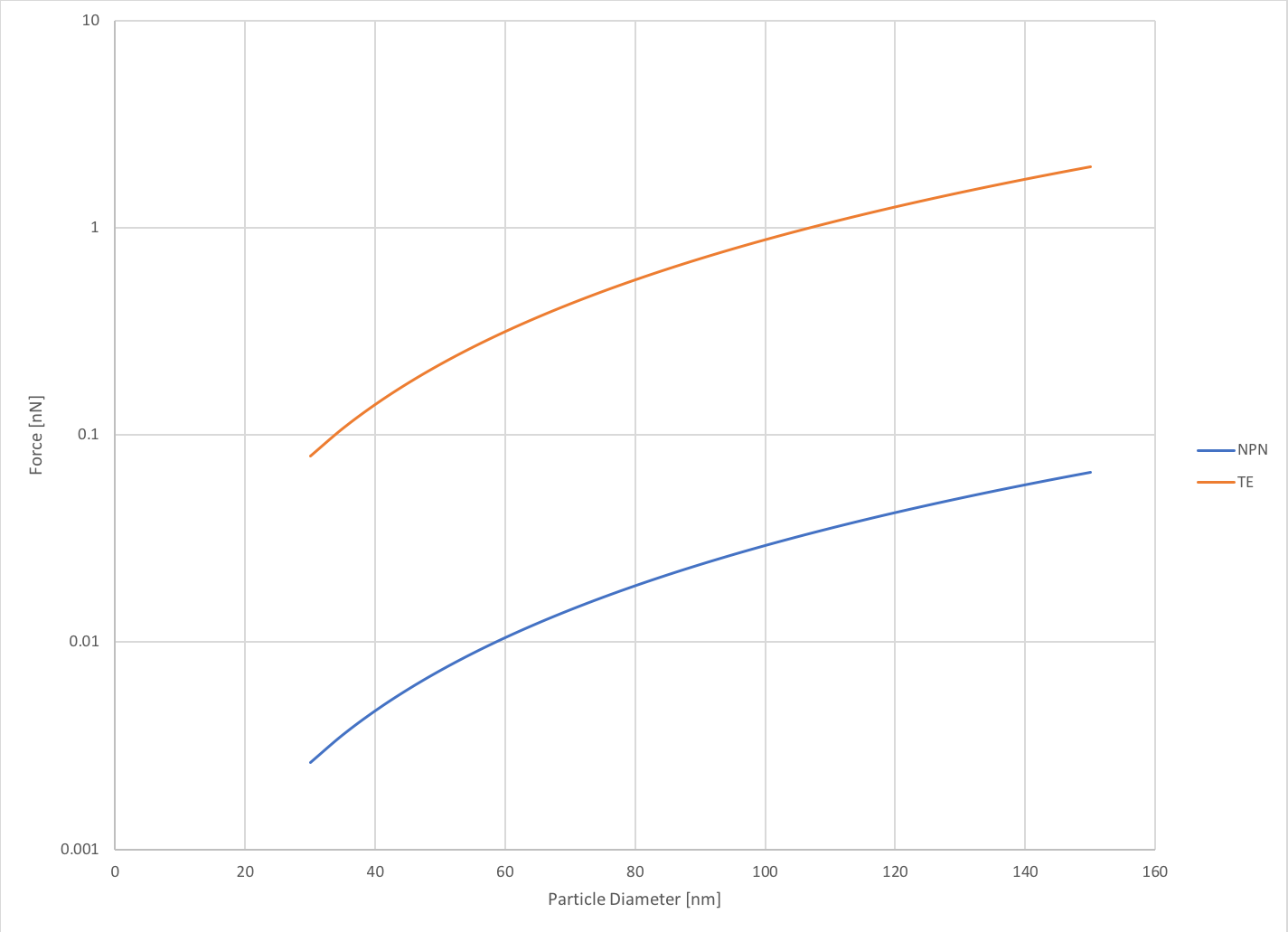
where [math] \Delta P [/math] is the transmembrane pressure, and [math] A [/math] is the cross-sectional area of the exosome. To gauge the forces experienced on particles within the size distribution of exosomes, I varied the diameter between 30-150 nm. I took the pressure value from my previous post, where I determined the transmembrane pressure experimentally. Additionally, I explored how the exosomes would behave in a track-etch membrane system under much higher pressure. As we can see below, the forces experienced on exosomes in the NPN system are quite small (0.003-0.07 nN) but when we model the TE system the forces are much higher (0.08 – 2 nN). This data is plotted on a semi-log scale for better visualization of the comparison.
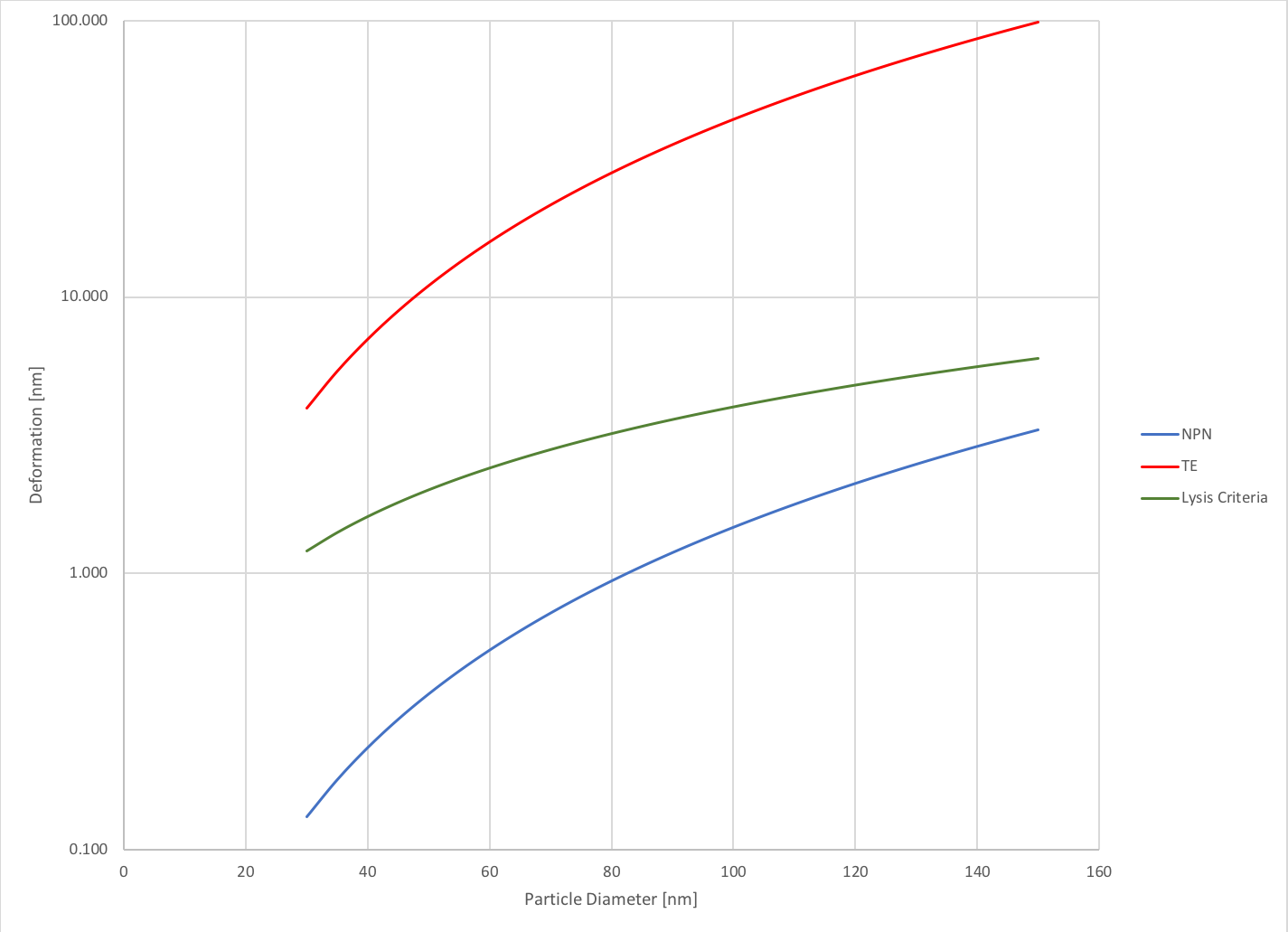
Now, these forces may seem very small, but we can use the earlier determined stiffness and calculate the deformation experienced by these vesicles under these forces. I plotted this data as a function of particle diameter, which allows us to see how the size of the particle affects the deformation. As expected, the larger the particle, the more it deforms. Furthermore, the particles deform significantly more in the TE system than they do in the NPN system. The exosomes deform up to 2.2% of their original size (for 150 nm exosomes) in the NPN system while in the TE system they deform up to 66% (for 150 nm exosomes). This data is also plotted with a criteria for lysis that Rick told me about. He said that a change of 4% is considered considered a condition upon which cells lyse and a condition that we should avoid. As we can see, the NPN captured exosomes do not approach this 4% criteria, but the TE vesicles all experience changes well above this threshold. This data is further indicative of the shortcomings of TE membranes for capturing exosomes. The data is again plotted on a semi-log scale.
While this data shows that exosomes are still likely nowhere close in mechanical properties to gold nanoparticles (I mean kind of obviously), they are still rigid enough in our systems that approximating them as rigid particles for the purpose of modeling should be accurate enough that we don’t have to consider the magnitude of the deformability in our models. Furthermore, the data presented in this paper is for liposomes, which are simple lipid bilayers. They do not contain the complex structure of exosomes that would include proteins in the membrane, which have been postulated to increase the stiffness of the membrane. This data would be best confirmed with AFM measurements, which I might take a day sometime in the future and add on to this.