Collagen IV is deposited below human umbilical vein endothelial cells
Introduction
My previous post focused on attempts to reproducibly generate collagen IV with human umbilical vein endothelial cells (HUVECs) so that we would be able to establish protocols for basement membrane protein detection and degradation measurement. Following my presentation on November 18 2020, there were some questions regarding whether or not the observed collagen IV was being produced on the apical side (above) or on the basolateral side (below) of the cells. I was also unsure of the answer since I had previously observed collagen IV deposition at various z-heights (Fig. 1). In order to generate an appropriate model of the endothelial basement membrane, collagen IV should be localized below the cells which would correspond to the region between the endothelium and the outer part of the blood vessel in vivo [1]. I began by trying to remove the cells while keeping the underlying proteins intact. Trypsin, which would normally be used to remove adherent cells, was inappropriate for this task since it demonstrates non-specific protein digestion [2]. Instead, I examined ice-cold PBS and water as potential options for cell removal. Additionally, I took z-stacks of fluorescently-tagged collagen IV and nuclei and examined where each structure came into focus to determine their relative positions.

Methods
Cell culture
HUVECs were obtained at passage 1 from Lonza (cat. no. C2519A) and grown in EGM-2 media + Bulletkit (Lonza, cat no. CC-3162) + 1% penicillin/streptomycin at 37 °C and 5% CO2. Media was changed every 2-3 days. The cells were used up to passage 7. For the different tests, cells were seeded at approximately ~3×10^4 cells/well in 24-well polystyrene plates (VWR, cat. no. 82050-892) and were maintained under the same culture conditions.
Cell removal with ice-cold reagents
HUVECs which were seeded at passage 7 were maintained in 24-well polystyrene plates for 9 days. Cells were washed once with room temperature PBS prior to addition of either ice-cold PBS or ice-cold double-distilled water. The ice-cold reagents were applied as two 5 min washes. Following this, the cells were fixed with 3.7% formaldehyde for 15 min. They were then permeabilized with 0.1% Triton X-100 for 3 min before addition of anti-collagen IV antibody (1042) alexa fluor 647 (eBioscience, cat. no. 51-9871-82) diluted 1:20 in PBS and subsequent incubation for 2 hr at room temperature. Nuclei were stained with DAPI (ThermoFisher, cat. no. D1306) diluted 1:400 in deionized water for 3 min. A 1% formaldehyde storage solution was added to the cells prior to imaging. All microscopy was performed on a Leica DMI6000B with a Q-imaging Rolera EM-C2™ EMCCD Camera. A positive control consisted of cells that were exposed to TrypLE™ Express Enzyme (1X), no phenol red (Gibco, cat. no. 12-604-013) for 4 hr prior to fixation, and a negative control consisted of cells that were not treated with any attempts at dissociation.
Z-stacks with deconvolution
HUVECs which were seeded at passage 5 were maintained in 24-well polystyrene plates for 11 days. Cells were washed once with PBS before incubation with anti-collagen IV antibody (1042) alexa fluor 647 diluted 1:400 in EGM-2 media + Bulletkit + 1% penicillin/streptomycin at 37 °C and 5% CO2 for 2 hr. After another PBS wash, a 500nM solution of SiR-DNA nuclear stain (Cytoskeleton, Inc. cat. no. CY-SC007) mixed in EGM-2 media + Bulletkit + 1% penicillin/streptomycin was added and the plate was returned to the incubator for 1 hr. The plate was imaged on an Olympus IX81 Inverted Fluorescence Microscope, and 9 µm high z-stacks were taken at 40x magnification with a 0.6 µm step size between images. The Olympus cellSens software was used to run a deconvolution algorithm on the z-stacks in order to reduce blurring from out-of-focus regions [3].
Results
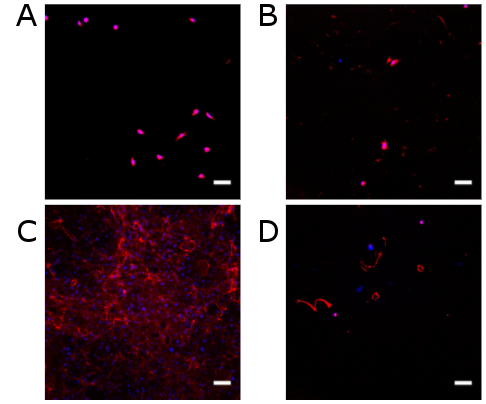
Ice-cold PBS and ice-cold double-distilled water were both effective at removing cells (Fig. 2A,B). However, they also removed collagen IV. Water left behind a little more collagen IV compared to PBS (Fig. 2B), but this hardly matched the deposits present when cells were not removed (Fig. 2C). PBS and water were comparable to treatment with TrypLE for 4 hr (Fig. 2D).
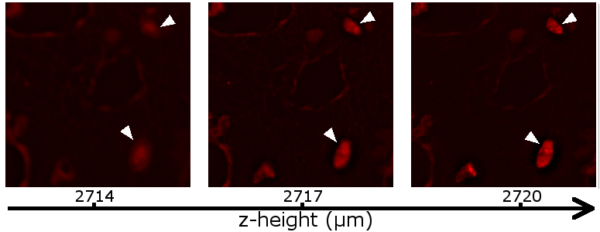
For the z-stacks (Fig. 3), SiR-DNA was chosen to stain the nuclei because it appears in the same fluorescence channel as the anti-collagen IV alexa fluor 647 stain. An alternative approach could have involved staining the nuclei with DAPI, but this might potentially influence height determination due to the chromatic aberration associated with all lenses [4]. Following deconvolution, the z-stacks were manually inspected to determine where the collagen IV and nuclei were in focus. In every examined stack (8 images taken across 4 wells), collagen IV appeared to be at a lower height than the nuclei (Fig. 4).

Discussion
The treatments with ice-cold PBS and double-distilled water were very effective at removing HUVECs from the 24-well plate. They also failed to leave substantial amounts of collagen IV behind for analysis. At first, this appeared to provide evidence that collagen IV might have been localized on the apical side of the cell, with the exception of the small deposits left behind following the ice-cold water wash. However, it was also feasible that collagen IV was located below the cells and was stripped away due to tight adherence to the cells. The z-stacks obtained with the Olympus microscope seem to provide some evidence for this possibility.
I originally intended to collect z-stacks with Rochester Institute of Technology’s confocal microscope in order to create high quality 3D fluorescence images, but realized too late that this microscope does not have a compatible holder for multiwell plates. In the future, it should be feasible to repeat this experiment on the confocal in order to confirm the results. It is worth noting that my observations do not account for the fact that I have observed collagen IV deposition at different z-heights as mentioned in the introduction, so this is worth investigating further. Importantly, this work was carried out in 24-well plates rather than the µSiM device that will be used for my future investigations into inflammation’s effect on basement membrane proteins. Transitioning to the µSiM will provide another opportunity to investigate where collagen IV is being deposited by the cells in a different culture environment.
References
[1] C. Leclech, C. F. Natale, and A. I. Barakat, “The basement membrane as a structured surface – role in vascular health and disease,” Journal of Cell Science, vol. 133, p. jcs239889, 2020.
[2] J. T. T. Baird and C. S. Craik, “Chapter 575 – Trypsin,” in Handbook of Proteolytic Enzymes (Third Edition), N. D. Rawlings and G. Salvesen, Eds., ed: Academic Press, 2013, pp. 2594-2600.
[3] R. Hard, J. Hipp, M. A. Tangrea, and J. E. Tomaszewski, “Applications of Image Science in Pathology and Cell Biology,” in Pathobiology of Human Disease, L. M. McManus and R. N. Mitchell, Eds., ed San Diego: Academic Press, 2014, pp. 3723-3759.
[4] K. W. Dunn and E. Wang, “Optical Aberrations and Objective Choice in Multicolor Confocal Microscopy,” BioTechniques, vol. 28, pp. 542-550, 2000/03/01 2000.