DNA Permeability of SiMPore Filters is Tunable by Changing Ionic Strength
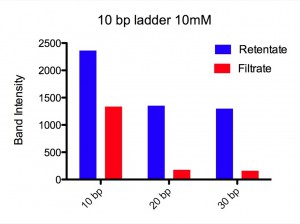
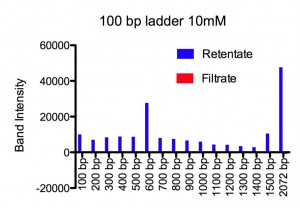
Our setup copied Jess’s well in a Teflon block. The chips (SCO21) sat over the well filled with 50 ml buffered (pH ~ 6.0) NaCl solution. On top of the well Paul placed a 20 ml drop of the same solution with DNA ladder added. Two types of ladder were used, referred to as a 10 bp ladder (contains 10 bp to 330 bp in 10 bp steps) and a 100 bp ladder (contains 100 bp to 1500 bp in 100 bp steps and a 2072 bp band). Each of these was run at two different NaCl concentrations, 10 mM and 155 mM. After 24 h at RT, Paul ran the retentate and filtrate on agrose gels, and integrated the band intensity for each DNA length.
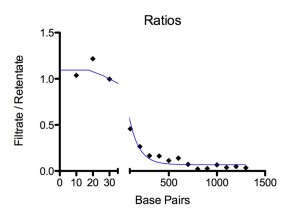
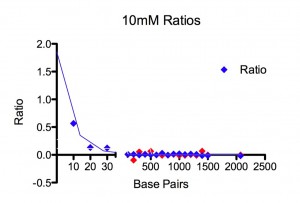
Results. For 155 mM Paul found all lengths, from 10 bp to 2070 bp, at detectable amounts in the filtrate. There was a plateau for 10-30 bp at ~ 100 % permeability followed by an exponential drop that reaches a permeability of 1/e at 100 bp. For 10 mM, using the same two ladders, he could only detect 10-30 bp in the filtrate. No DNA from 40 bp to 2072 bp could be seen on the gel for the filtrate. An exponential fit to the three data points, 10, 20, and 30 bp, gives a permeability of 1/e at 8 bp. Figures and fitting parameters are shown below.
Discussion. We don’t yet know the pore size distribution for the SCO21 chip but they are very thin and, if they are like the SCO24 chip, the large pores are 30-33 nm and some of these are merged into bigger pores. DNA length is 0.34 nm/bp. So a 100mer is 34 nm long. Short DNA is very stiff. A 20mer is a stiff rod. A 100mer is a flexible rod. At ~500 bp DNA will start to wrap back on itself. Simplistically, one might expect the dynamic radius of a 100mer to be ~15 nm and, if the maximum pore radius is 15 nm, permeability should drop around 100mer. Maybe our finding is fortuitous, but that is what we see at 155 mM NaCl. At 10 mM NaCl, the membrane charge becomes (a lot?) more negative and the DNA slightly more negative. The electrostatically constricted pore behaves as though it has a radius of ~ 5 nm.
Conclusions. Our methods need to be improved and subsequent experiments conducted with higher detection sensitivity. We are excited. What do you think?


Bill and Paul – We’ve been dreaming about separations this tight forever. The difference between 10 bp and 30 bp in 10 mM is ~6.5 nm !?! Very exciting indeed. We do worry about how the quality of such separations depend on time. Repeating this out at different time points would be a nice experiment for us if you can afford to do it.
This is double stranded DNA, right? What is the diameter? There is a nice physical model of separation to develop here. Have you done 0 M salt or is there a reason you can’t?
The really large stuff (~400 nm?) getting through in 155 mM seems like a breach in the membrane, or it is snaking its way through. The stuff is floppy out here I know, but still.
Of course now that you’ve made a post you are obligated to present it. There is no NRG meeting next Tuesday, but there is the week after. Feb 24 from 9-11AM.
It is just awesome to have you aboard. Thanks!
Very interesting work – thank you for the post!!
Do you happen to know how low you can drop the salt concentration before the double-stranded DNA denatures? Is there a limit on how high the salt background can be?
Since DNA (especially double-stranded) is so linear, we may see 2 transport modes. When the “effective” pore size is larger than the length of the molecule, I assume transport will be normal and comparable to globular molecules. However, when the pore size drops below the DNA length there must be the possibility for some level of aligned transport or “snaking” as Jim and Jess describe it. Perhaps some of this is visible in the 155 mM ladder? Does the exponential fit hold out to the longest molecules, or does it seem to flatten out at a non-zero level? I’m not exactly sure how this transport would behave, but the dependence on the molecular length would probably relax a little – once you approach/enter the pore, sliding through seems easier, so length would be less important. Also I would expect the dynamics to be much different in our short pores, than in the very long pores. In our case, a long molecule would be simultaneously entering and exiting for a period of time.
Neat stuff! I look forward to hearing more… Thanks!
Nice work!
Pore distributions can be found here. All images are in the photo gallery, and here’s a link to wafer SC 021. There are definitely some elongated pores in this membrane that have an equivalent diameter up to around 30nm.
Thanks for the enthusiastic responses
Re Jim’s Qs
Time allowed for diffusion is a variable we will have to monitor. All we have done so far is to run thymidine on the SC021 membrane for 2 and 24 h. At 2 h, it was still far from equilibrium; at 24 h, the concentration was equal on both sides.
Yes the DNA is in a double helix conformation. Diameter = 2 nm.
We have not done 0 M salt but hope to soon.
The large stuff! Yes, we are guessing that there are some big pores that let the high molecular weight stuff through. In our very first experiment, done last summer we saw high permeability to a 10mer and zero to a 2500 bp supercoiled plasmid; that experiment used UV (260 nm) to measure DNA concentration, which is more sensitive than the gels Paul ran to get the data we showed you.
Paul and I will come to the NRG meeting on the 24th. Where is it?
Re Chris’ Qs
The temperature DNA denatures at depends on the CG content and MW. We are assuming that our DNA ladders were ~50% CG. At high MW and 10 mM NaCl, denaturation is expected between 65 and 75 deg C. At 10 bp, 50% CG, 10 mM, I guesstimate denaturation should be in the 20-30 deg C range. So for high MW, we can probably go down to the 2 mM NaCl range at RT and still have native duplex but for 10 bp, we were probably already on the edge in the above 10 mM NaCl experiment. We can determine whether DNA is denatured or not but have not yet done so.
In the other direction, DNA is super stable at high salt; 6 M LiCl can be used.
“snaking” might happen. We need to talk with some of the molecular dynamics experts. I have feeling this would not be terribly hard to simulate with some the new computational packages. And my polymer physical chemistry is not good enough to predict whether or not translation diffusion is fast compared to rotational diffusion and how to partition rotation around the helix axis from rotation perpendicular to the helix axis.
The exponential fit at 155 mM NaCl reaches a non-zero (well above detection threshold) plateau somewhere between 500 and 1000 bp. Because we had complete exclusion for the plasmid DNA mentioned above, I suspect this had to do with a small number of large pores and that the plateau level would have increased with time.
We are also wondering what happens when one end of the “needle” finds a pore. My first guess is that if the needle is long compared to the pore diameter, the dynamics of rotation is more likely to pull it back out than drive it through. I suspect the polymer chemical physicists have worked this out decades ago.
Jess
Thanks for the pointers. In looking at the photos, it seems to me that there are merged pores that could be over 50 nm long. With pores in that size range, the big DNA may make it through.
Plans
Jess gave us 6 more SC021 membranes. We hope to have a new set of blocks with 20 micro liter wells this Wednesday. That will double our sensitivity. We are ready to ramp up and start running experiments in parallel. Lots to do. Variables of interest: time, temperature, ionic strength, and double vs. single stranded DNA. What would help our rate of progress now is more membranes, and if possible ones with pore size maxing out in the 20-25 nm range and free of larger merged pores. Don’t know if that is possible yet?
In our histograms we show the Equivalent Diameters, or basically a diameter given a circle with the area of the pore. It’s true that some of these pores have a longer dimension, so here’s a look at the major axis: Some pores go out as far as 60nm in one dimension.
Thanks Jess for looking at asymmetric pore sizes. I think these are going to be important given the large asymmetry in DNA.
BUT the link didn’t work. I got a “object not found!” error. Can you try it some other way?
Our research project is aimed at determining the physical-chemical mechanisms by which ionizing radiation (e.g., X-rays, gamma-rays) damage DNA. One important class of damage is single strand breaks; if these occur on opposite strands within 10 base pairs (bp) of each other, a double strand break occurs. Associated with every break is the release of the DNA base (Cyt, Gua, Ade, or Thy) at that site. Using Sympore filters, we believe it will be possible to quantify strand breaks and free base release in new ways and at higher sensitivity.