Initial MgF2 Nanomembrane Cytocompatibility Study (Live/Dead)
I want to verify that I can grow a confluent monolayer of cells on my MgF2 nanomembranes. I have tried to do this in the past, but have only gotten sparse cells overall before the membranes ruptured. I believe that part of the problem was that I was not seeding at a very high density of cells to start my experiment.
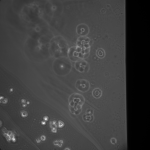
I used my MgF2 nanomembrane process to create 8 Flow-chip Format MgF2 nanomembranes (Wafer 1153, 50 nm thickness, 35 nm, 15%). The overall yield was 8/17 chips, almost 5o%. I used 4 of these chips in an experiment, placing them face up in a 6 well plate suspended over 1 mm high, 3 mm wide channels, allowing media to make contact to both sides of the nanomembrane. I used 4 1153 NPN chips as a control. The chips were attached to gaskets isolating the top membrane area from each other (simple stiction). The gasket assembly was sterilized with autoclaving prior to use. The NPN chips and the MgF2 chips were placed in different wells (4 chips per well). I used MDB131 Complete media to hydrate each channel, then I seeded P9 HUVECS on the flat surface of each chip at fairly high density (1:10 split of T-25 confluent flask, 20 uL/well). After letting the cells adhere for 1 hr, I flooded each well with 5 mL of media, and placed into the incubator. Visual inspection after 1 hr, prior to flooding, showed that 3/4 of the membranes were intact in each well and the cells were starting to spread out on all surfaces.
(One chip was broken in the NPN well, and 2 were broken in the MgF2 Well, B3)
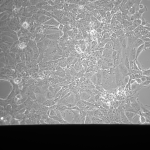
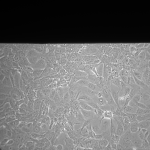
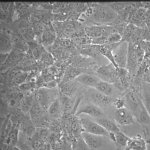
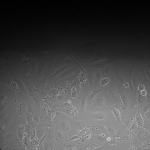
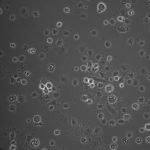
NPN Controls (24 hrs):
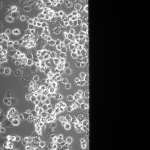
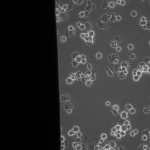
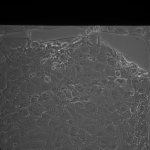
MgF2 Nanomembranes (24 hrs):
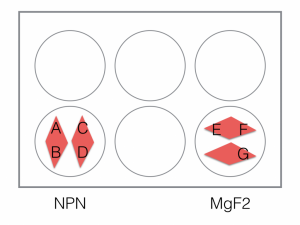
This was my first attempt to have really large area nanomembranes work in cell culture. Unfortunately, the additional steps in my cell culture process were not enough to make them survive even 24 hrs, though they did survive the wetting steps. I have had some success with smaller membranes (SEPCON format). With these items in mind, I ran the experiment again (HUVECs P11), trying to make the wetting as perfect as possible by hand. I also seeded at half the density I used before (10 uL of a 1:10 split). A, B, E, F received cells. C, D, G, did not.
Images are all 20x, except where noted
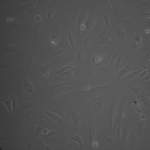
NPN Controls (2hrs):
MgF2 (2 hrs):
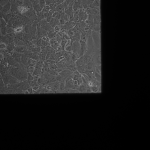
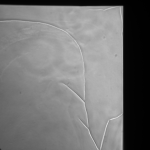
MgF2 (24 hrs):
G is not dissolving in cell media appreciably in 24hrs, preserving the cracks. I believe the fluid motion is exacerbating cracks in the MgF2, furthering small tears if they appear.


After observing these images, I attempted Live/Dead Staining. I prepared 10 mL of a 4 uM EthD-1 and 2 uM Calcein solution in PBS, vortexing to ensure even mixing. I aspirated the media from the wells, and replaced it with the Live/Dead solution, pipetting in 20 uL of solution into the channels and device wells before flooding the entire well with the remaining volume (5 mL total per well). I incubated the plate for 45 minutes. I swapped out the filters by hand manually, controlling the shutter for the Hg-lamp as well by hand.
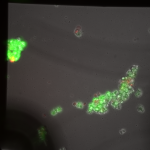
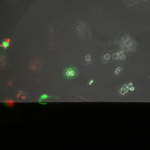
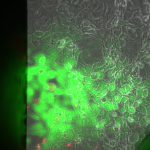
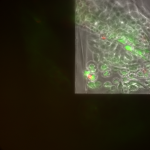
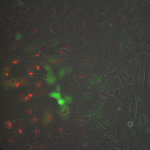
Live/Dead images (26 hrs):
The MgF2 cells appear to be mostly alive and not dead, with some red nuclei appearing. The source was not well aligned to the microscope; I was unable to make it better in the time alotted. I then made a bad choice to kill my cells with bleach (4 mL added to each well, 5 minutes of dwell time).
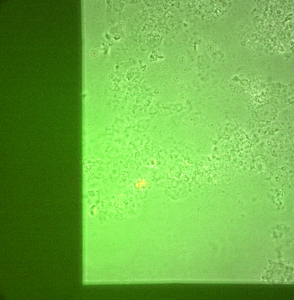

Overall, I have shown that I can have a large number of cells adhere to a large area of my MgF2 membranes, spreading and growing. No additional surface treatments were used in this experiment, save UV-ozone to attach gaskets. They are still very fragile, so multiple wetting steps are difficult to work with by hand. In the future, I will be using smaller windows to lessen the chances of breaking.
My next experiment will be to use pnc-Si capped NPN substrates as a control for cell attachment + growth (Josh Winans’ paper), and compare it to my MgF2 membranes. I plan to fix with Formaldehyde instead of bleach to keep the cells more intact next time.