Followup on biosensor project
This is a followup to my previous post, which can be found here.
Previously I reported that fluorescein diphosphate (FDP) incubated on the surface of an NPN chip would adhere well enough to be visible on the membrane after gentle washing and treatment with a fluorescence-activating enzyme (alkaline phosphatase, AP.) I also reported that pre-incubation with poly-L-lysine (PLL) blocked this fluorescence, hypothesizing that this was due to PLL binding too tightly to FDP and blocking access to it by AP.
While these results were promising, further experiments over the past week have led me to conclude that they were coincidental or indicative of poor experimental procedure, due to my inability to replicate them. Of the dozen or so chips I’ve attempted to adhere FDP to since my first experiment, only a handful have succeeded, and disparate results were often obtained from chips with identical treatments.
Jim and I have hypothesized that the trouble may be the salt concentration during the incubation step. Because the adherence we’re looking for should occur ionically, high salts could shield the opposite charges from one another and block adherences. Today, I repeated the original experiment with a new procedure, outlined briefly here:
- Isolate 8 chips from the wafer. Incubate half with 100 uL of 10 uM FDP in DI water, and the other half with the same volume of 10 uM FDP + 1 uM PLL in DI, for 90 min.
- Remove the incubating solution and rinse each chip three times with 100 uL DI water.
- Add to 2 chips from each half 20 uL of 0.08 ug/mL AP in tris buffered saline (TBS; 50 mM tris, 150 mM NaCl, pH 7.4) and add only 20 uL TBS to the other 2.
- Record fluorescein signal using FITC excitation laser.
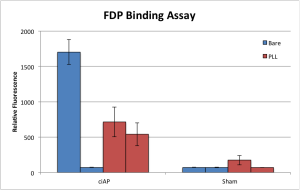
Something perhaps not immediately obvious is that I’ve doubled the concentrations of FDP and AP used in this experiment over the original — this is my way of trying to force the experiment to go the way I want, since all we’re interested in right now is getting it to work, before we invest too much time in it. The results this time are a little more comfortable than they have been historically:
Each bar here represents a single chip, with the fluorescence averaged over two separate windows. Of the two chips which did not receive PLL but did receive AP (here labeled “ciAP,” calf intestinal alkaline phosphatase,) one was very bright, while the other was indistinguishable from the bare-sham chips. This is exactly as I’ve come to expect of chips treated with just FDP: when it works, it works dramatically; but it usually doesn’t work. On the other hand, the PLL-treated chips were never as bright, but at least in this experiment they were much better behaved.
These results are in line with my understanding of the physics at play here. PLL is strongly positive and should bind tightly to the negative FDP molecules. While this should increase the amount of FDP hanging around on the chip after washing (since PLL sticks to the chip surface very well,) it should also reduce the availability of that FDP to its activator, AP. Thus, we see reduced signal when compared to the one bare chip which FDP stuck to, but the FDP did stick, apparently in 3/4 cases as opposed to 1/4 without PLL. Progress…?
Moving forward, I’ll try to repeat these results while also demonstrating the effect of ligating the AP to a large complex, as we intend to do for the biosensor proof-of-concept experiment.
The sham experiments did not see AP, right? If I understand that correctly, then PLL is working on much of the FDP but some of the FDP is not accessible as you say. There is still a nice jump from sham.
You would be better off with consistency and lower signal from PLL than inconsistency and higher signal without it. The all-or-none issue is a strange problem, but if PLL lets you avoid debugging it, lets use PLL.
Repeat this exact study first to see if we can avoid introducing voodoo.