Journal Club: Residual matrix from different separation techniques impacts exosome biological activity
Residual matrix from different separation techniques impacts exosome biological activity (PDF)
Lucia Paolini, Andrea Zendrini, Giuseppe Di Noto, Sara Busatto, Elisabetta Lottini, Annalisa Radeghieri, Alessandra Dossi, Andrea Caneschi, Doris Ricotta & Paolo Bergese.
Scientific Reports 6, Article number: 23550 (2016) doi:10.1038/srep23550
Abstract
Exosomes are gaining a prominent role in research due to their intriguing biology and several therapeutic opportunities. However, their accurate purification from body fluids and detailed physicochemical characterization remain open issues. We isolated exosomes from serum of patients with Multiple Myeloma by four of the most popular purification methods and assessed the presence of residual contaminants in the preparations through an ad hoc combination of biochemical and biophysical techniques – including Western Blot, colloidal nanoplasmonics, atomic force microscopy (AFM) and scanning helium ion microscopy (HIM). The preparations obtained by iodixanol and sucrose gradients were highly pure. To the contrary, those achieved with limited processing (serial centrifugation or one step precipitation kit) resulted contaminated by a residual matrix, embedding the exosomes. The contaminated preparations showed lower ability to induce NfkB nuclear translocation in endothelial cells with respect to the pure ones, probably because the matrix prevents the interaction and fusion of the exosomes with the cell membrane. These findings suggest that exosome preparation purity must be carefully assessed since it may interfere with exosome biological activity. Contaminants can be reliably probed only by an integrated characterization approach aimed at both the molecular and the colloidal length scales
Summary from today’s journal club to be added later…
The main take home point from this paper:
Exosome purification techniques that utilize precipitation agents (PEG, etc.) leave a residue that seems to prevent these exosomes from interacting with cells in future experiments/procedures. (This is bad for therapeutic exosome use)
In the introduction the authors also make note of several reports of immune-protein complexes/aggregates that can contaminate exosome purifications and affect proteomic and transcriptomic studies. (This is bad for diagnostic exosome use)
Statements from the paper to expand upon during the discussion:
“Autoimmune diseases, hematologic disorders, infections, and cancer associated with elevated exosomes counts are also characterized by accelerated formation of immune and protein complexes11. These aggregates share several biophysical parameters with exosomes – such as size, surface charge and light absorption – which may strongly affect vesicles purication11,17,19. Protein contamination can also alter or even invalidate proteomic and transcriptomic studies on exosomal proteins and genetic material20–23”
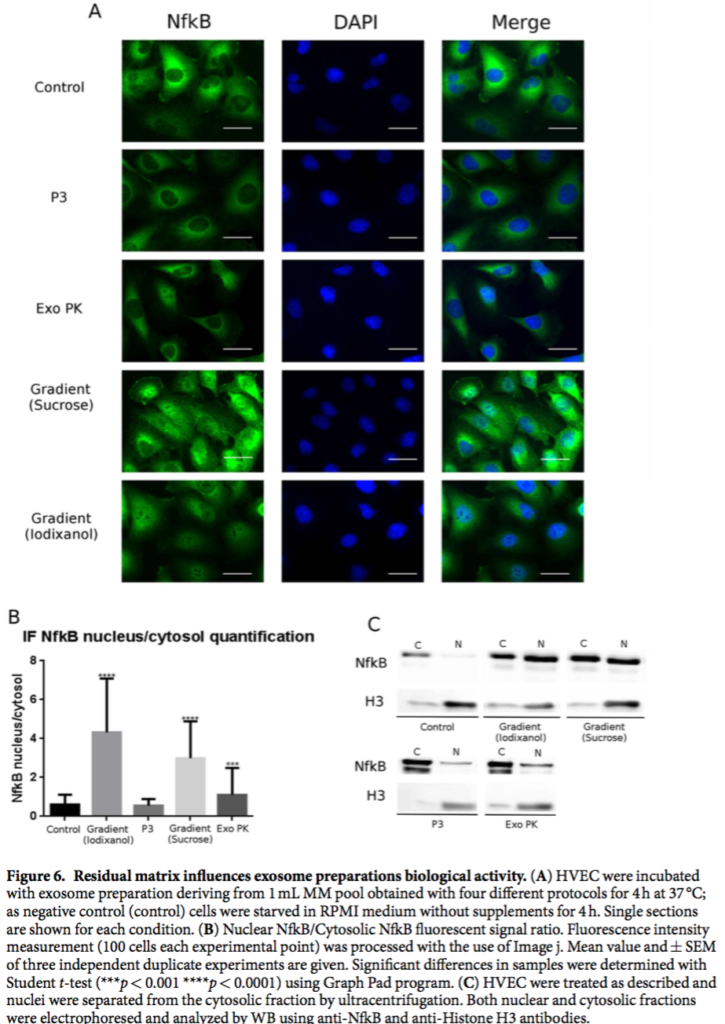
“From the sample images reported in Fig. 6 we learn that cells incubated with the P3 preparation showed no signicant differences with respect to those treated with the control buffer; cells treated with the Exo PK preparation showed a faint NfkB nuclear signal. In stark contrast, cells incubated with the pure exosome prepa- rations obtained from iodixanol and sucrose gradient, showed a clear strong NFkB nuclear translocation signal (Fig. 6A,B). ese data were also con rmed by WB analysis of nuclear extracts reported in Fig. 6C, where an intense NfkB signal in the nuclei of cells incubated with gradient preparations is evident.
The origin of these results is probably related to the two steps mechanism, which determines the interaction between exosomes and target cells. The initial step is binding of exosomes to the target cell surface through ligand-receptor recognition and lipid-mediated interaction27. Numerous membrane-associated proteins decorate exosomes membranes and are functionally active on target cells35,36, driving exosome binding within minutes37,38. In the second step the attached exosomes are internalized by endocytosis pathways, which differ depending upon the recipient cell type39. In view of this mechanism and of the fact that in the contaminated samples the exosomes are embedded or surrounded by an exogenous matrix, we can reasonably infer that in the P3 and Exo PK preparations the residual matrix interferes/hampers the interaction between exosomes and cell membranes, and in turn their internalization and activation of NfkB translocation. Alternatively (or in concomitance) it is possible that residual matrix contains proteins that can block NfkB activation, particularly in P3 preparation.”
Other related papers:
Immunoglobulin free light chains and GAGs mediate multiple myeloma extracellular vesicles uptake and secondary NfκB nuclear translocation (PDF – This is the authors prior paper looking at Multiple Myeloma exosomes and HUVECs)
Comprehensive Proteomic Analysis ofMesenchymal Stem Cell Exosomes RevealsModulation of Angiogenesis via NuclearFactor-KappaB Signaling (PDF – This is a related paper by different authors that was just published. They showed that MSC exosomes induce angiogenesis through NfkB signaling)