ASEMV Conference Review
Last week, I attended the American Society of Exosomes and Microvesicles (ASEMV) conference in Monterey, Ca and presented a poster on the work that I’ve been doing. Despite a very low turnout for my poster session (due to the scheduling of a long break before and dinner after), I met with some very interesting people and learned a lot of things. I’ve included a summary of the talks that I took notes on over the two days that I was there.
Monday
Tom Gingeras: CHSL, Tailoring of Extracellular RNAs. This was an interesting talk, mostly about exosomal RNA content. That is kind of important as we want to be able to know which RNAs are in the highest abundance in exosomes. Noted that many features of EVs (e.g. proteins and such) are found in ultracentrifugation supernatent, which brings in to question the method of isolation. The main take away was that the method of isolation significantly affects the results of analysis and that ultracentrifugation (UC) is not scaleable in terms of time and cost. This figure is from one of his papers about the RNA content of exosomes. Basically, I will have to refresh my knowledge of all types of RNA.
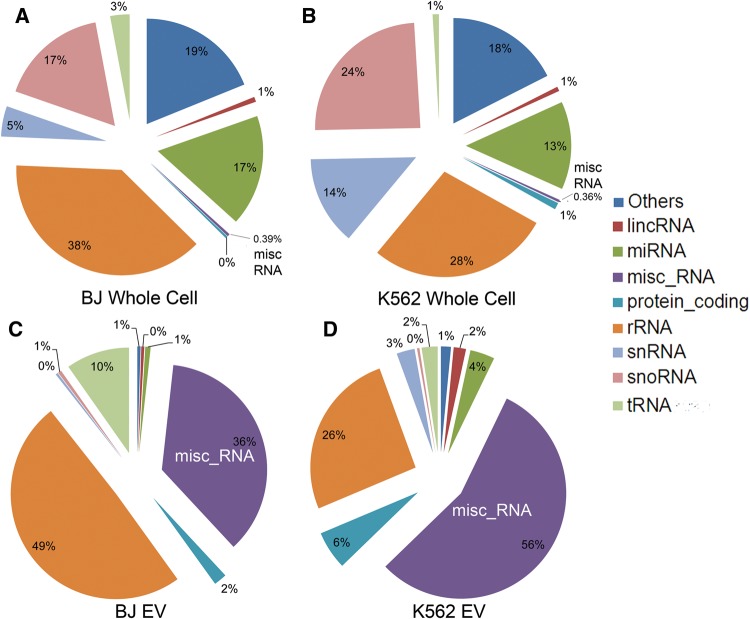
Harmeet Malhi: Mayo Clinic, STARD11-mediated Ceramide Transport is Necessary for Release of Lipotoxic Extracellular Vesicles. Didn’t take too much away from this other than her application was non-alcoholic steatohepatitis (NASH) and that the ceramide transport protein STARD11 is important for the transfer of ceramides in EVs (which is basically what the title says).
Rienk Nieuwland: University of Amsterdam, Prediction of venous thromboembolism in cancer patients. Didn’t really talk about this too much in terms of EVs. He showed a screen shot from the O magazine website, which I thought was funny. Basically, the exosomes in your saliva help to induce clotting, so sucking on a wound will help it to heal faster as cell derived vesicles have coagulant tissue factor when found in saliva. Talked about a review that he co-authored titled Methodological Guidelines to Study Extracellular Vesicles.
Shilpa Buch: UNMC, Exosomal miRNA-mediated loss of pericyte coverage at the blood-brain barrier: Implications for morphine-mediated potentiation of neurodegeneration. HIV associated neurological disorders. Noted that miRNA 9 increased in exosomes. miRNA 29b in exosomes leads to astrocyte apoptosis. Morphine reduces pericyte coverage, causes cognitive impairment. Showed CD63 and flotillin characterization of exosomes.
Kyungsun Choi: KAIST, Engineered exosomes loaded with super repressor IkB, a potent inhibitor of NF-kB pathway, for treatment of septic shock syndrome. Technology is called EXPLOR. Said that current techniques of loading exosomes only load 2% of the total product and are passive loading techniques. They want an active loading technique. Use optogenetics. Yim N et al. Nature Comm. 2016. CIBN-tetraspanin. Cover protein on molecule in exosome activates and deactivates with blue light. The loaded molecule is associated with the cover protein. Exosomes were used for delivery of CRE to mice. Deliver tumor suppressor switch to exosomes, exosomes effectively cleared to lysosomes by phagocytic pathway. srIkB loaded exosomes increased LPS injected mice survival. 80% after 80 hours vs. 0% for untreated after 40 hours.
Florencia Menay: CEFYBO-CONICET, Exosomes isolated from ascites of T-cell lymphoma-bearing mice induce tumor specific immune response. This talk was very difficult to understand and she mostly presented her isolation and characterization, which wasn’t any different than conventional methods. She did show that her exosomes generated an immune response, which is kind of a debated topic for exosomes. She did show that there was 1.7 x 10^8 exosomes/mL isolated from ascite media.
Jing-Hung Wang: Stanford, High affinity anti-HER2 scFv-directed extracellular vesicle-mediated mRNA-based gene delivery inhibits growth of HER2-overexpressing human breast cancer xenografts by prodrug activation. Plasmid labeling of HER2 to fluoresce. pXPORT loading of exosomes. (I can’t really find information about this whole pXPORT thing, I think it’s unpublished work.)
Joshua Hood: University of Louisville, Cyclical Electrical Field Flow Fractionation or Melanoma Exosomes. This was an interesting concept, but not very memorable. Their data wasn’t really convincing and basically they were using electrophoretic properties to control exosome separation. I think the most important thing that I learned fro this talk was that exosomes have a negative collodial stability.
Swetha Srinivasan: Abbvie, Utilizing extracellular vesicles from patient plasma to identify biomarkers of autoimmune disease. Didn’t really present too much about specific biomarkers. I took away that they used 5 donors for biological replicates and had 5 technical replicates from each donor. They then pooled donors and added an artificial miRNA that they could search for. Their biological variation was higher than their technical variation, but they did still have significant technical variation indicating that this is a tricky process to repeat. I also learned that the Clontech RNA detection kit requires 1 ng of tRNA.
Gareth Willis: Boston Children’s Hospital, MSC Exosomes Ameliorate Experimental Bronchopulmonary Dysplasia and Restore Lung Function Through Macrophage Immunomodulation. British fella with looks that every girl likely desired. Kinda arrogant, he was a doctor not really a scientist. They isolated exosomes fromMSC conditioned media. They used TFF to assist with their separation (didn’t say with what, couldn’t find him to ask afterwards) and they “concentrated 50 x using TFF” and then ultracentrifuged their sample afterwards (using an iodixanol gradient).
Travis Antes: Cedars-Sinai Med Center, New targeting methods that manipulate the surface of EVs for enhanced tissue delivery. Cardiosphere derived exosomes. Used a technology for uptake of labeled exosomes using ‘cloak’. Can’t find too much about this cloak, unpublished work right now.
Marsh Durban: ReNeuron Ltd., Neural Stem Cell-derived exosomes induce senescence and apoptosis in cancer cell lines. They produce exosomes commercially. Process 25-50 L (yes, that is correct, they process liters of material) of conditioned media for exosome purification. They use hollow fiber TFF, but didn’t elaborate too much on that. When they release their captured exosomes, they get 10^11/10^12 particles (EVs?)/mL. The check for sterility by checking mycoplasma and endotoxins, they check size and concentration using the NanoSight and they check ID and functionality using a “potency” assay. Potency is perhaps the better choice of words rather than viability.
Jan Lötvall: University of Gothenburg, Diversity of extracellular vesicles – implications for biomarker discovery. All EV isolates contain multiple EV populations. Davide Zabeo, JEV 2017. They look for exosomes from tumors directly, not from blood or plasma. They use a high pH/high salt treatment to disrupt the membranes to do membrane proteomics. They showed that within the exosome membrane there were plasma membrane proteins, endoplasmic reticulum (ER) membrane proteins and mitochondrial inner membrane proteins. They also cited a website that has a database of exosome proteins and microRNAs called EVpedia. I created my own login to be able to access the database, it’s completely free.
Siyang Zheng: Penn State, Rapid isolation of extracellular vesicles using lipid nanoprobes. Estimate a 15 minute processing time. Their goal is cancer diagnostics, checking for metastasis. They have previously done work with ctDNA, CTCs (for 10 years) and exosomes (for 3 years, about the same amount of time that we have been doing it). They use frozen plasma and a “few hundred μL of blood plasma”. It was pretty funny, he seemed to forget his audience and gave a whole talk about the background of exosomes to a bunch of exosome experts. In fact, that was a pretty common theme for people who it was obvious was their first time and they didn’t make the presentation for their specific audience (they just changed the title slide). Their technology is a lipid labeling probe that works as follows:
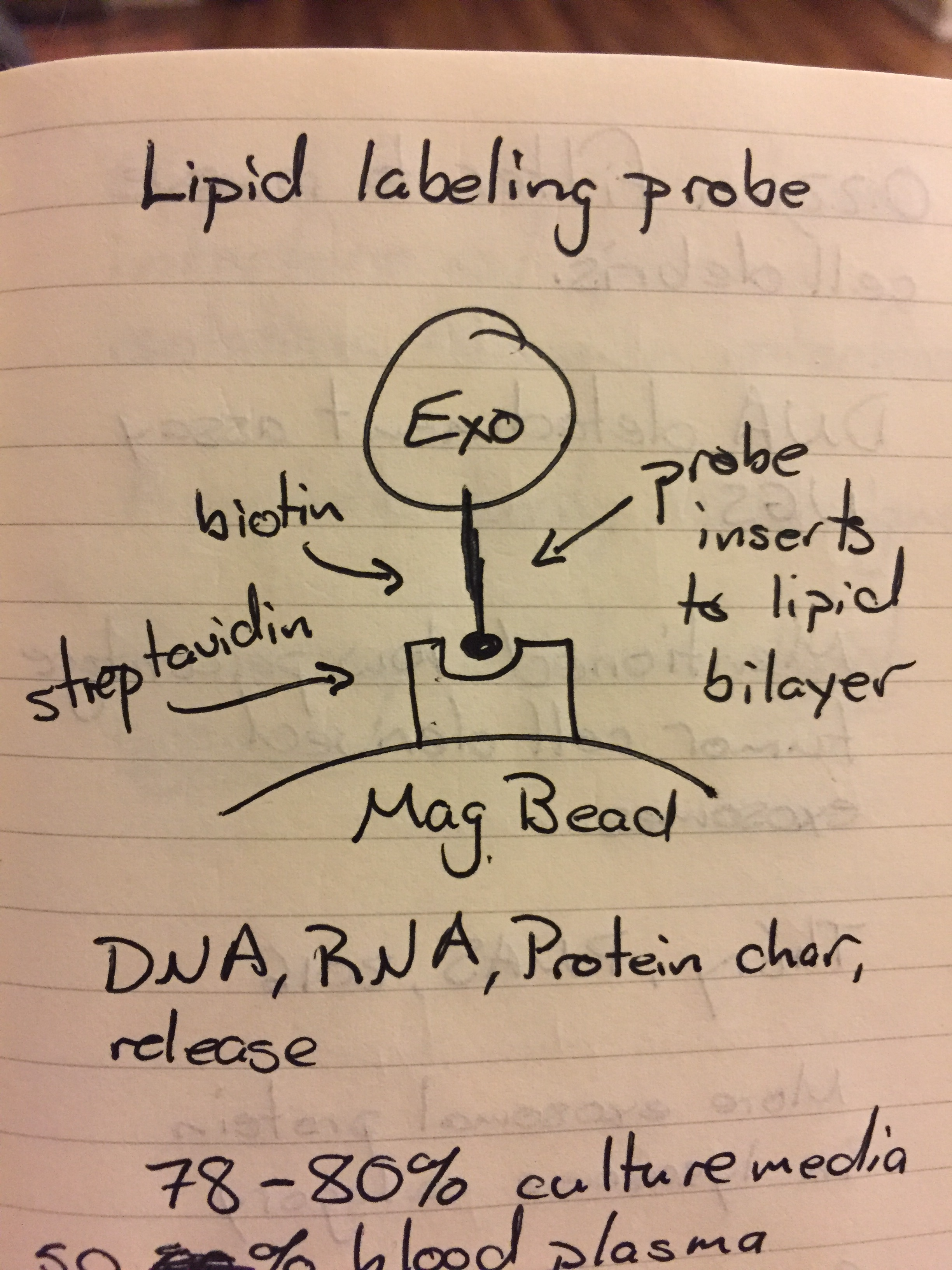
They can do DNA, RNA and protein characterization and then they can release the exosomes from the beads (but not the probe). They get about 70-80% recovery from cell culture media and 50% recovery from 1:3 diluted (note that it is diluted) plasma. They do cryo-SEM of their EVs on and off the probe but don’t use a label meaning that these could literally be anything! They also do a 0.22 μm prefiltration step, which could result in lost vesicles and people brought up the fact (during questioning) that in plasma, there are many lipid binding proteins (albumin!) that could interfere with this capture technique. Also, because of the lack of a label, they could be getting a false positive from apoptotic bodies as this technique captures any vesicle with a lipid membrane!!!
George Daaboul: NanoView Biosciences, High-Throughput Characterization of Single Exosomes and Extracellular Vesicles. They can identify particles ranging from 40 nm to several microns in size. They can get concentration, size and phenotype. They can work directly from plasma and use small volumes (5-100 μL). They use affinity based capture. Their technology is bilayered Si-SiO2 to enhance the optical signal to visualize items below optical wavelengths using a simple microscope. This is a plot of their contrast curve; basically the larger the vesicle, the better the contrast:

They haven’t really correlated their results to EV sizes, but they took Zika (d = 70 nm..?) and were able to image in SEM and using their system to correlate the exact sizes that their system generated. They don’t give a processing time, but upon asking afterwards he told me that their flow is passive (no pumping) and that to get the best results they have to let it sit overnight. They also use 1:3 diluted plasma and can get activated platelet contamination.
Tuesday
Alain Brisson: University of Bordeaux, Methods of imaging, quantification and isolation of exosomes/EVs. This was the keynote talk and perhaps my favorite one of the conference. First off, this guy was a French version of Sir Ian McKellen, which was pretty freaking awesome. He was also super nice and was pretty interested in what I was doing. Most of the people there got their EM images from a core, but he does all his own imaging, so he understands the process and the data very well and doesn’t just show lousy EM images. He uses immuno-cryo-TEM and flow cytometry to characterize the EVs. Also resistive pulse sensing to get size information and stuff. He notes that there is a single method for detection of EVs, that we must use a combination of methods. He also notes that there are no good methods for isolation as many are very non-specific for the populations that we are after. His first question was whether or not it was possible to image EVs in pure, unprocessed plasma using cryo-TEM and they showed that they could in fact do this. Seriously check out the paper that I linked there, there are some phenomenal images of EVs. He noted that using the common negative staining for exosomes in TEM is really messy for pure plasma and thus it is best to use cryo-TEM (it did win a Nobel prize a couple of weeks ago, so there’s that…). He notes that cryo-TEM is the gold standard for EV imaging because we can image them in their native hydrated state. This means that as soon as we get the cryo attachment for our TEM, I will be on it to check and see how our samples look! He gives a lot of good papers (Arraud, J Thromb & Hemo, 2014, 12:614, Brisson, Platelets, 2017 (I need ILL to get this one), Lötvall, JEV 2015 (they study ejaculates for their exosomes, no seriously! ?????). They study phosphatidylserine (PS) exposing EVs. They also show that a minority of EVs have Anx5 binding (so not that many are platelet derived). SINGLE STEP LABELING IS CRUCIAL (NOT MULTISTEP!!!)!!!!! That was an extremely important point that might help with our labeling. They even double immunogold labeling (two targets). The size of EVs are determined by their cargo. Koifman J. Struct Bio, 2017. Cloutier EMBO Mol. Med. 2013(2) 235-249. Melo Nature 2015 523: 177-185. Show that Glypican-1 is specific for pancreatic cancer specific exosomes. Brisson Cytometry A, 2016. This paper is really interesting as it discusses labeled EVs (so EVs that have some sort of marker) and what it shows is that the counts are actually really low (10^4/10^5 range)! They also show that not all exosomes have CD63!
Suresh Mathivanan: La Trobe University, Exosomes from bovine mile reduces the tumor burden and attenuates cancer cachexia. Zhang, Cell Research, 2012 (plant miRNA found to be transferred to human cells). Hypothesis: Milk increases cancer progression. Ca stabilizes exosomes (CaCl2). Results: Milk EVs decrease tumor volume/burden and proliferation. Milk exosomes have anti-cancer effects. However, they also increase metastasis with the primary tumor intact (without the primary tumor, they decrease metastasis).
David Lubman: University of Michigan (sorry Dean), Proteomic Analysis of Exosomes from Patient Serum Samples. They use ultracentrifugation to isolate their exosomes but run it 5 TIMES (!!!) to remove albumin. They note that their yields are really low. They then run mass spec on exosomes for CD63, CD9, TSG-101 (use LC-MS/MS).
Antonio De Maio: UCSD, Isolation of glycoprotein characterization of human urine extracellular vesicles. Uses ZnSO4 before freezing to preserve exosomes and help to remove uromodulin (Tamm-Horsfall protein) from the urine. Gave a protein marker that is highly specific to exosomes derived from the bladder or the kidney only (nowhere else in the body): galectin 3 binding protein (G3-BP). Also showed that for patients in septic shock, the exosomes varied quite significantly in size while in healthy patients, the exosomes were very consistent in size. This was shown across multiple patients, not just one patient multiple times.
All in all, this was a very good conference to attend and perhaps once we publish, they will invite us for a talk next year. They also talked about the ISEV 2018 conference next year and the Gordon Conference on Extracellular Vesicles in Maine next year. Sorry for the long post, but hopefully it has good info for you guys!