Separation of Extracellular Vesicles using Ultracentrifuge
Hi everyone,
One of the objectives in the last aim of my PhD is to investigate the effect of soluble factors secreted by adipose derived stem cells (ADSCs) on wound recovery of fibroblasts (FBs). We have categorized these soluble factors to 3 subpopulations including: large-EVs, small-EVs, and non-EV associated molecules. For the last 3 weeks, i have been running experiments to separate these subpopulations from each other and to optimize this protocol for further experiments. This post is more about the principles for ultracentrifuge and some results from my latest experiments.
Here is the protocol recommended by Terry et, al,. for isolation of exosomes by ultracentrifuge which has been widely used in the field:
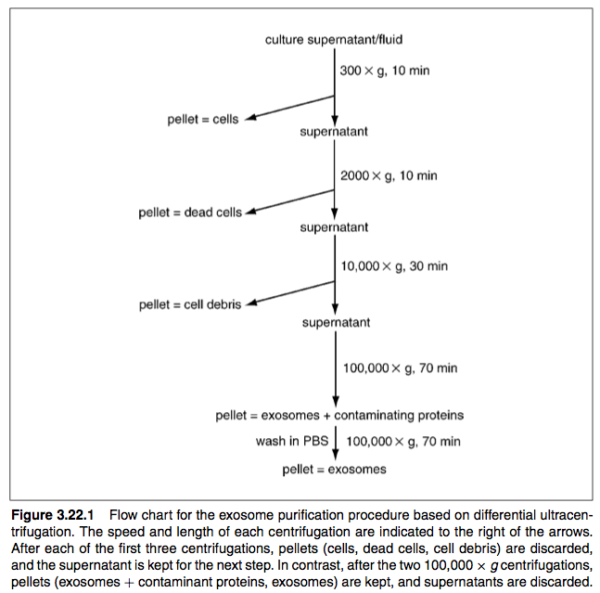

The protocol includes multiple spinning step in order to separate different size particles, the third snipping step is suggested to be used for pelleting cell debris. However, this step can be used for micro vesicles as well or (large EVs). It is important to note that we do this step for 30 minutes at 15000 g. Particles have different sedimentation coefficient depending on their size, mass and the viscosity of their suspension media and this can be calculated by equation 3 (Based on Stocks law).
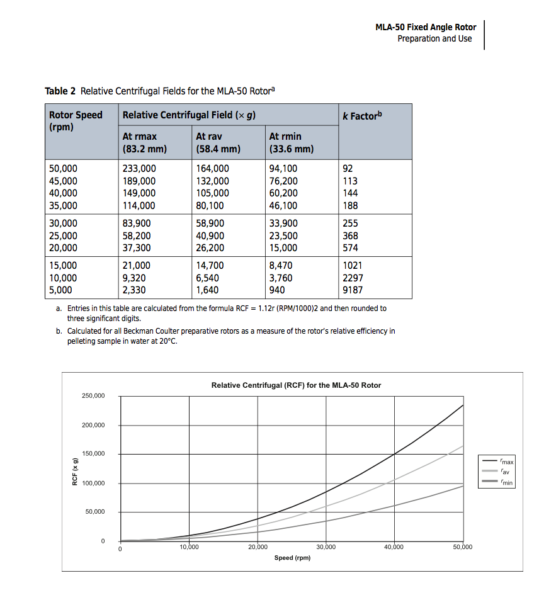
In the old EV papers, unfortunately people did not report information about their rotors properly. It’s important to report information about the rotors so that other people can replicate the same set of experiment. This is actually recommended in Minimal studies for studying extracellular vesicles 2018 and i was personally told this multiple times from Tom and Jim. This is why we need to optimize the protocol to our rotors based on the chosen references.
Every rotor has a cleaning factor (K factor) at it maximum speed which can be calculated as following. If the experiment is run with lower speed that maximum, the K factor can be calculated from equation 2. The lower the K-factor, the better the rotor. Equation 4 shows the relation between the K-factor, time of centrifugation and the speed. Equation 5 shows how equation 4 can be used to convert a centrifugal run with one rotor so that an equivalent run can be applied for another run. In our lab, we have 2 types of rotor and i’ve optimized our systems based on these 2 rotors.
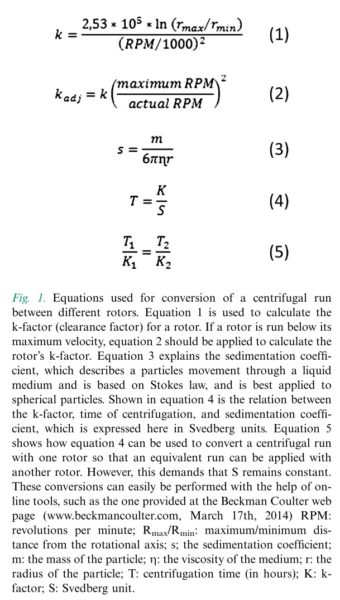

In addition to that, for fixed angle rotors, depending on the location of the particles they experience different g forces. Particles close to the bottom of the tube experience the highest g. So this also needs to be paid attention to when we are optimizing our samples.

So now i am focused on optimizing the protocols for us to be able to isolate different size particles with ultracentrifuge. So far, the yield seems to be around 5-25 % which is actually also reported by others. In order to optimize the system, speed and time are two factors to play with. I decided to test if i can separate larger EVs from smaller EVs and i tested two possible ways of doing: 30 minutes for 15000 RPM and 8 minutes for 50000 RPM.
30 minutes for 15000 g:

8 minutes for 50000 g:

Then i decided to isolate the EVs derived from ADSCs with 2 hr of spinning at 50000 RPM. This experiment was done with 5 mL of conditioned media which is promising and shows that we might be able to isolate EVs with conditioned media from T25 flasks.
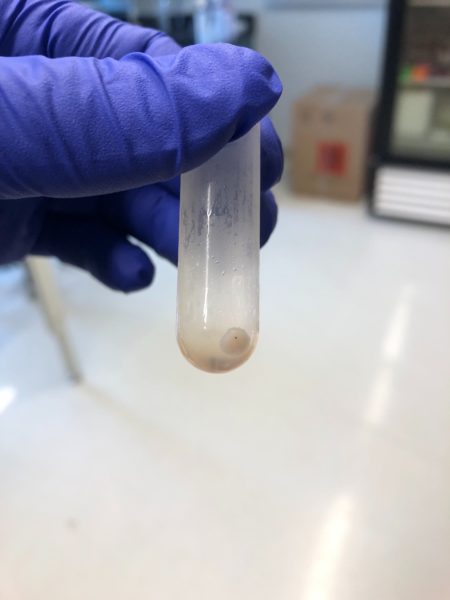
I am planning to optimize the system to have an EV free supernatant and a pellet with all EVs. One argument is that we will end up co-isolation of non-EV molecules and this can be characterized using western blot and EM. My next step is to achieve the EV-free supernatant and check the surface markers in order to test the purity. I tested this once but i didn’t have much luck with i, however I also tested 10% FBS and tried to spin down for 12 hours and the supernatant was not fully particles free, however the pellet was clear and visible at the bottom of the tube. This means that we can make our own EV-free media for the future experiments.
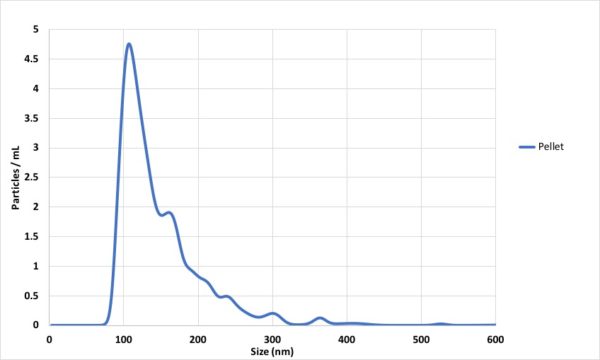
At the end, NTA has 2 factors to play with which are camera level and detection threshold to optimize the measurement. i find it very interesting that i now i can visually tell if the particles are typical EVs, proteins or large EVs based on their scattering pattern and their motion. based on my experiments with Hansa exosomes, ADSC-EVs and T24-EVs, camera level of 13 and detection threshold of 5 are the best ones and i always try to keep this constant between my measurements.
ADSC pellet (2 hr 50000 g) 2019-07-23 18-25-18 (Converted)
ADSC pellet (30 min 15000 g) 2 2019-07-23 16-29-58 (Converted)
Thanks