ECM alignment mechanosensing and cell polarization using microporous membranes as masks
Objective: Determine the minimum required area and number of exploratory cytoplasmic protrusions for sensing ECM alignment and causing cell polarization. Hypothesis: Cells only need to sense collagen alignment through exploratory protrusions extended during initial phases of cell seeding and not throughout the entirety of the cell, and a minimum number or size of protrusions is required for the cells to sense alignment. Rationale: Cell polarization has been observed to be sensitive to the degree of alignment in collagen gels. By using microporous parylene membranes, the area of aligned ECM a cell is exposed will vary, depending on the porosity, as seen in the figure below. By changing the average number of pores within the vicinity of a cell, the number and size of exploratory protrusions can also be controlled, giving insights into the minimum area and number of projections required to polarize cells on an aligned substrate. Approach: Using microporous parylene membranes as masks, we will control the number of sites at which protrusions can sense ECM alignment and also the area over which exploratory protrusions can sense ECM alignment.
Using our knowledge from our own PhD projects, Adeel (Heterogenous collagen alignment), me (versatile microporous membranes) and Zahra (Cell-substrate interactions) are working together to work on this project. Our rotation student (Mehran) is also providing some help with this project.

In order to test our hypothesis, we a device that puts aligned/unaligned collagen in full contact with parylene membranes, while providing the ability to align collagen using shear stress (Figure 2). After going through several device prototypes, we are now working with channels inside a PDMS mold, which is filled with collagen and the collagen aligned with shear stress and due to the surface treatment on the inside walls of PDMS channel, we are able to peel off the PDMS, without detaching the collagen, leaving three collagen strips with 200 um height. This method is not perfect, but we have several alternatives in mind which we are going to test for future experiments.
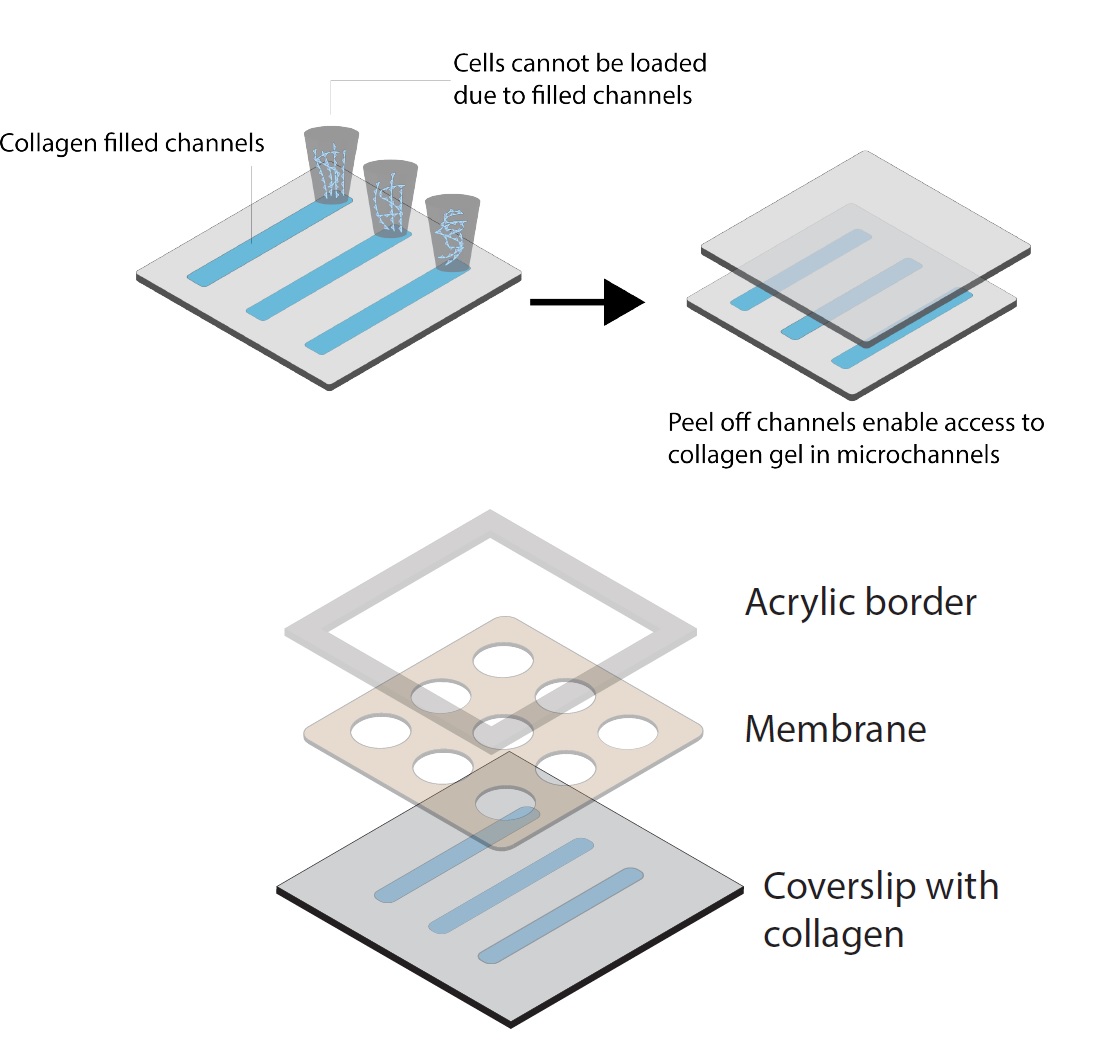
One concern we had initially, which turned out to be a legitimate one was that the height of the channel (200 um) as compared to the 1 um thickness of the membrane can lead to gap problems (data not shown here) and cells (at least in some areas) do not come into direct contact with collagen through the membrane. To overcome this, we came up the idea of attaching the membrane first to a acrylic top with a rectangular opening in the middle. Our rotation student also tried to do some work by surface modification of parylene membranes. The following video shows the successful outcome of these efforts (since Mehran didn’t have any controls for confocal imaging, we cant yet determine what led to good result (Video 1). Although the cell culture results indicate that even without surface modification and only by the change in device desgin, the gap issue was solved.

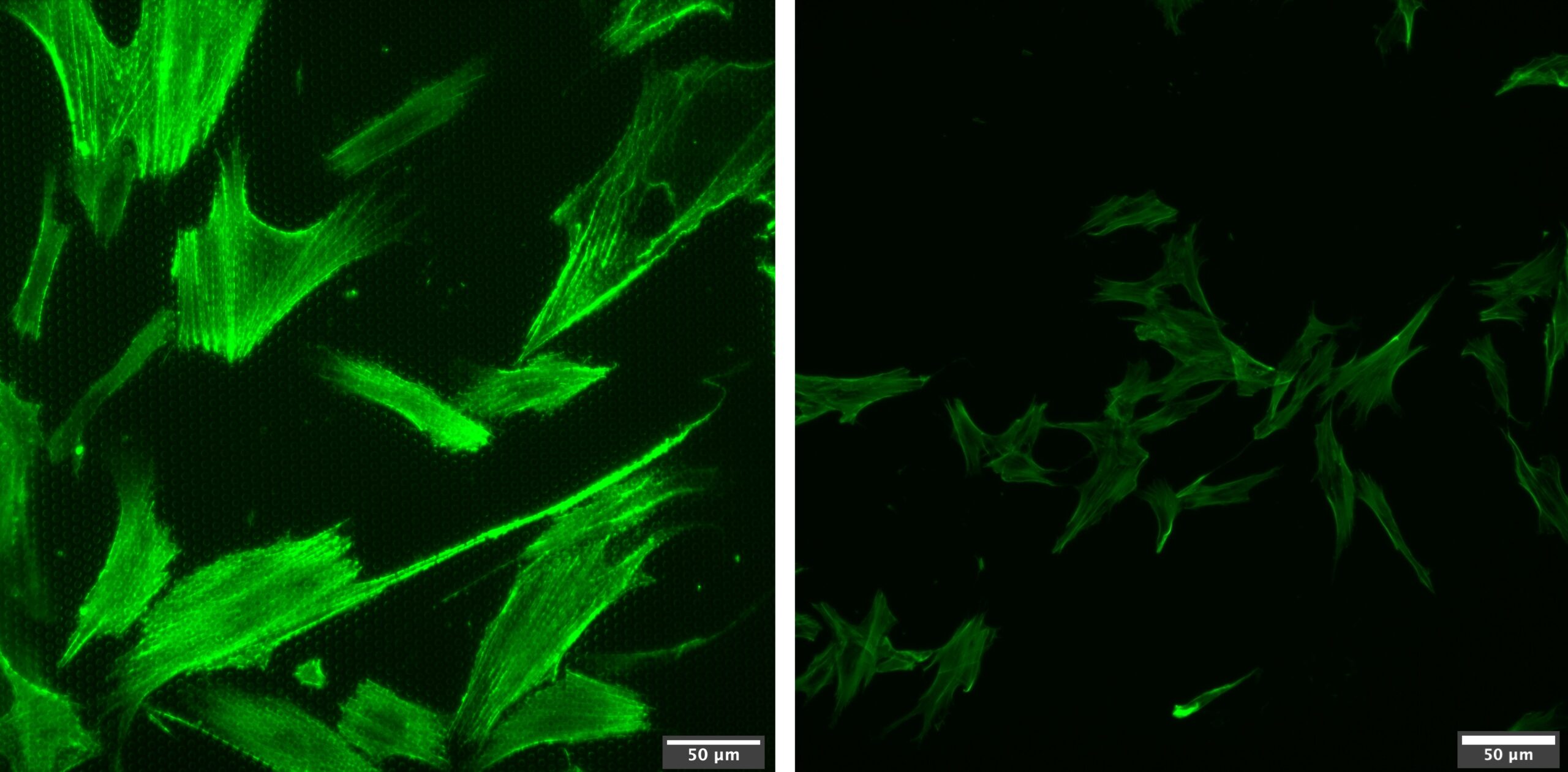
Although we will eventually may move back to cancer cell model, due to the problems Abhyankar MDA-MB231 cells had, we used ADSCs, which have proved to be much more robust cells for such preliminary studies. After a couple of issues with confocal microscopy and difficulties with seeding densities, we seeded the devices with enough low cell density to be able to fix and stain for cytoskeleton after 24 hrs and be able to observe pronounced cell ECM sensing in the absense of significant cell-cell communication. The following is a representative image of cell ECM sensing and cell polarization on 3 um porous membrane vs TCP control (Figure 3). What we observed from this study was that although cells were able to sense the collagen and make protrusions through the membrane, 3 um pore size was not enough for cells to aligning in the direction corresponding direction. For future work, we aim to tune membrane properties and pore geometries as well as optimizing the device design.
https://drive.google.com/open?id=1HNP37LJhOPo_lXetPTkjESGFi1csA4Pj