in situ Permeability Analytical Solution Summary – Unhindered vs Hindered D
Introduction
Pages regarding our in situ permeability assay development can be found in these links:
- Theoretical Underpinnings of Small Molecule Permeability Measurements in the µSiM (Part 1: Approach)
- Theoretical Underpinnings of Small Molecule Permeability Measurements in the µSiM (Part 2: Experimental Validation)
- Theoretical Underpinnings of Small Molecule Permeability Measurements in the µSiM (Part 3: Application to Cell Barriers)*
- ALine Modular Device: in situ Small Molecule Permeability (Confocal Microscope)
Briefly, this assay utilizes confocal microscopy to monitor passage of dye from the top well of a µSiM device to a set focal plane below the membrane (usually ~100 µm). For cell-seeded devices, permeability is calculated by fitting our experimental data to a constant flux equation (see Part 3 of Theoretical Underpinnings for details).

This equation is very sensitive to the diffusion coefficient (Dz) inputted for the fluorescent molecule. Since we see a significant hindrance to diffusion with our coated, cell-free devices compared to uncoated, cell-free devices, we wanted to test the relevance of each diffusion coefficient to analysing the cell-seeded in situ data. The goal was to determine if we need to account for the contribution of our coating solution to passage of the fluorescent molecule, in order to get an accurate calculation of endothelial, rather than system, permeability. Therefore, for each cell-seeded data set, we calculated permeability using an unhindered, free diffusion coefficient (found in the literature or calculated using Stokes-Einstein), as well as using a hindered, measured diffusion coefficient (measured using the in situ assay across coated, cell-free devices fit to a free-diffusion equation; see Part 2 of Theoretical Underpinnings).
Results
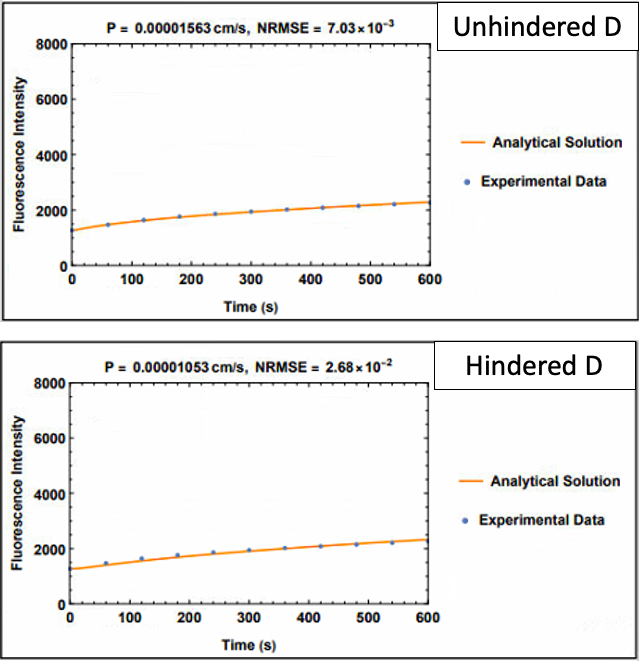
The plots above are comparative plots from lucifer yellow across an EECM-BMEC monolayer. Please note that findings are the same for hCMEC/D3 monolayers. While the analytical solution using the constant flux equation fits the experimental data well using both the unhindered diffusion coefficient (6.83e-6 cm2/s) and hindered diffusion coefficient (2.15e-6 cm2/s), the unhindered D solution fits better, with less error (NRMSE = normal root mean square error).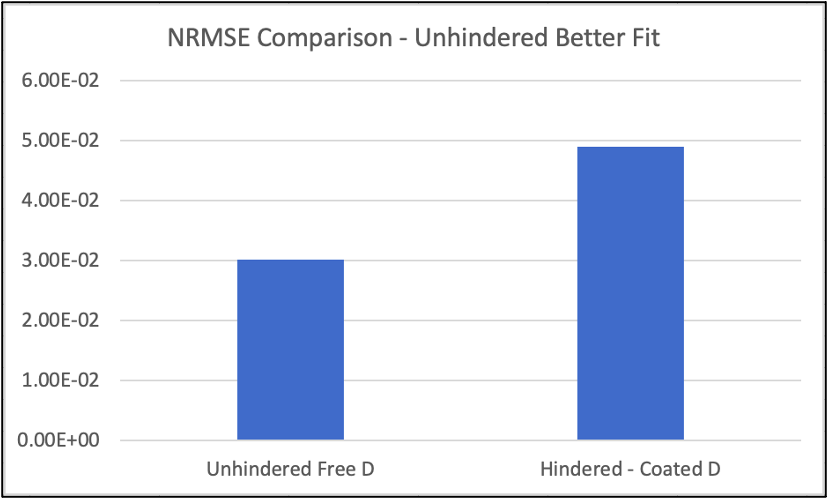
I repeated this for all data available for lucifer yellow across both hCMEC/D3 and EECM-BMEC monolayers (n=10 in total), and consistently found less error using the unhindered diffusion coefficient. This is an objective way of saying the unhindered D is the appropriate diffusion coefficient to use in the in situ data.
This finding was surprising, given the significant hindrance seen in our coated devices. However, to further drive this point home, we looked at a comparison of calculated hCMEC/D3 permeabilities between molecules of two different sizes (10 kDa dextran and 457 Da lucifer yellow) and compared the findings to data from our endpoint, sampling assay.
Interestingly, 10 kDa dextran is less sensitive to the coating solution, as it is hardly hindered across cell-free coated devices. Because of this, the hindered diffusion coefficient is very close to the unhindered value and the permeabilities are similar using both D. However, for lucifer yellow, only permeabilities calculated using our unhindered, free diffusion coefficient match permeabilities from our sampling method (2-way ANOVA, purple vs green bars, LY). Another important point is that we lose our ability to detect differences between molecules of different sizes when using the hindered D (pink bars, both molecules).
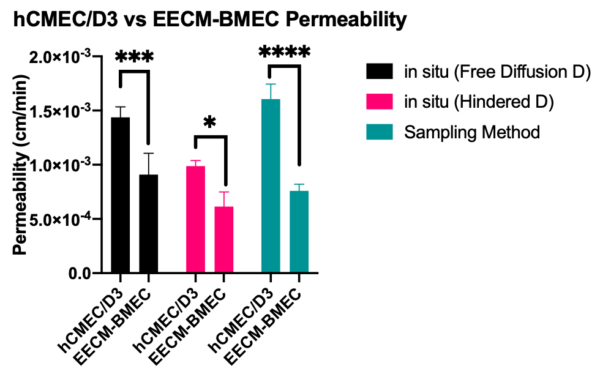
Finally, I compared barrier function between hCMEC/D3 (weaker barrier properties) to EECM-BMEC (tighter barrier properties). For all methods of analysis, we are able to detect significantly tighter barriers for the EECM-BMEC monolayers (n=3-4, 2-way ANOVA). Interestingly, for the tighter EECM-BMEC monolayer, there was no significant difference between any method of analysis. In other words, similar to 10 kDa dextran across hCMEC/D3, the smaller endothelial permeability is less sensitive to the analytical method (unhindered or hindered D). I should note the the NMRSE, which looks at goodness of fit of the analytical solution to the experimental data, as stated earlier, is still lower for unhindered D for the EECM-BMEC monolayers.
Limitation of unhindered, free diffusion coefficient?
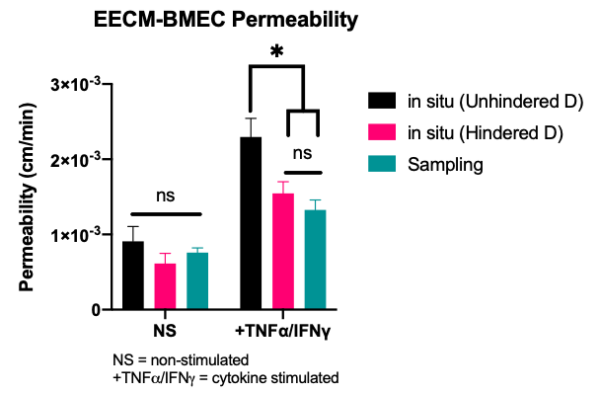
The only potential limitation to using the unhindered, free diffusion coefficient in the in situ permeability analysis, is when we treat our monolayers with cytokines (0.1 ng/ml TNFa + 0.1 ng/ml IFNg). Now we see no significant difference to our sampling data when we use our hindered D (pink vs green bars, TNF/INF), compared to a significant difference when we use the unhindered D (black vs green bars, TNF/INF)(2-way ANOVA).

Despite the better match to sampling data, the NRMSE is still lower (fit is better) for the unhindered D. However, as seen from the plots above, we actually lose the excellent fits seen with unstimulated cells. This indicates we may need a new equation to analyse stimulated monolayers. The constant flux equation may not be appropriate anymore.
Conclusions
Unhindered, free diffusion coefficients are appropriate for analysing endothelial permeability in the in situ permeability assay. There are three points to support this:
- Analytical solutions using Unhindered D fit experimental data better (lower NRMSE).
- Unhindered D permeabilities match sampling data for lucifer yellow, whereas hindered D solutions are significantly lower than sampling data.
- Unhindered D shows significant difference in permeability between 10 kDa dextran and smaller Lucifer yellow, whereas hindered D solutions do not.
We were able to detect significant differences in permeability between weaker hCMEC/D3 and tighter EECM-BMEC monolayers, regardless of the diffusion coefficient used.
The analytical solutions using the constant flux equation do not match well to the experimental data for cytokine-treated cells. We may need to find a new solution for this data.
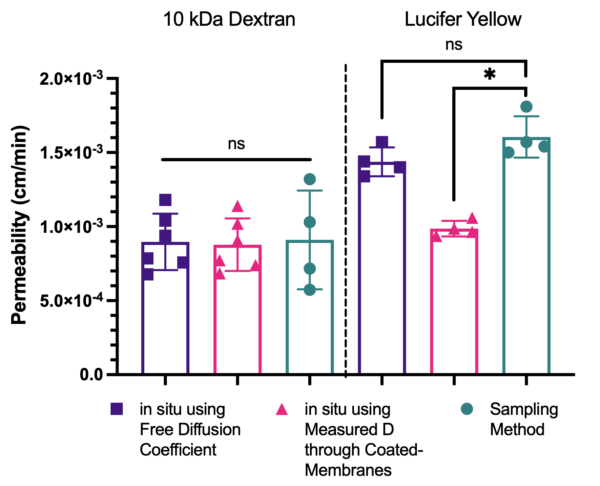

Can you please update this post with the concentrations of cytokines you used?