WEKA segmentation produces higher fidelity npMgF2 models
Summary
My analysis of npMgF2 has relied heavily on both tomogram and segmentation processes. The tomogram generation has thus far been sufficient, giving enough detail to our eye to identify different structures, but turning the tomograms into 3d models has been a lot more labor intensive. Using a neural network classifier (Weka, out of a New Zealand research lab, plugin found in Fiji), I am better able to show different grain structures within npMgF2.
Weka Classifier
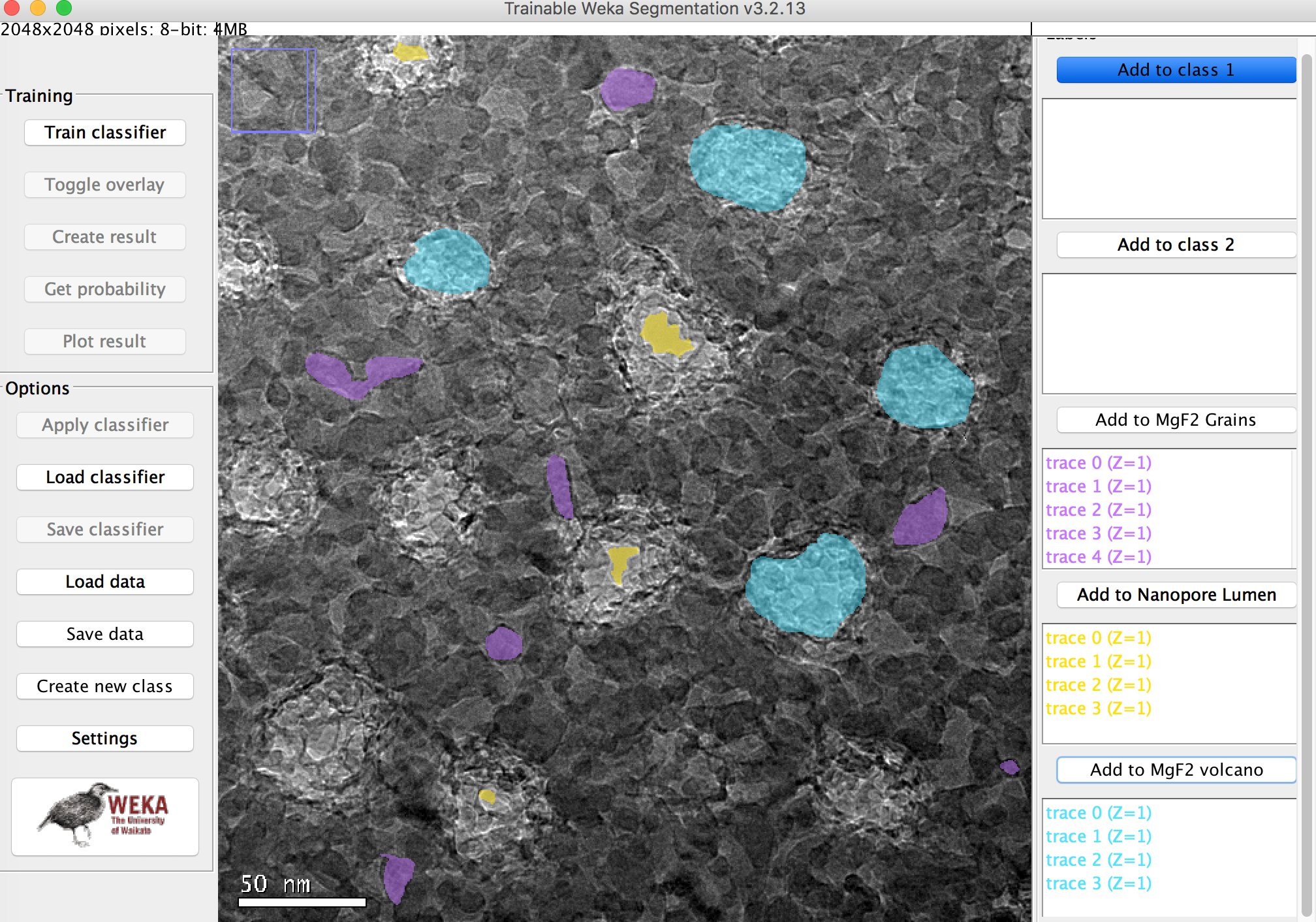
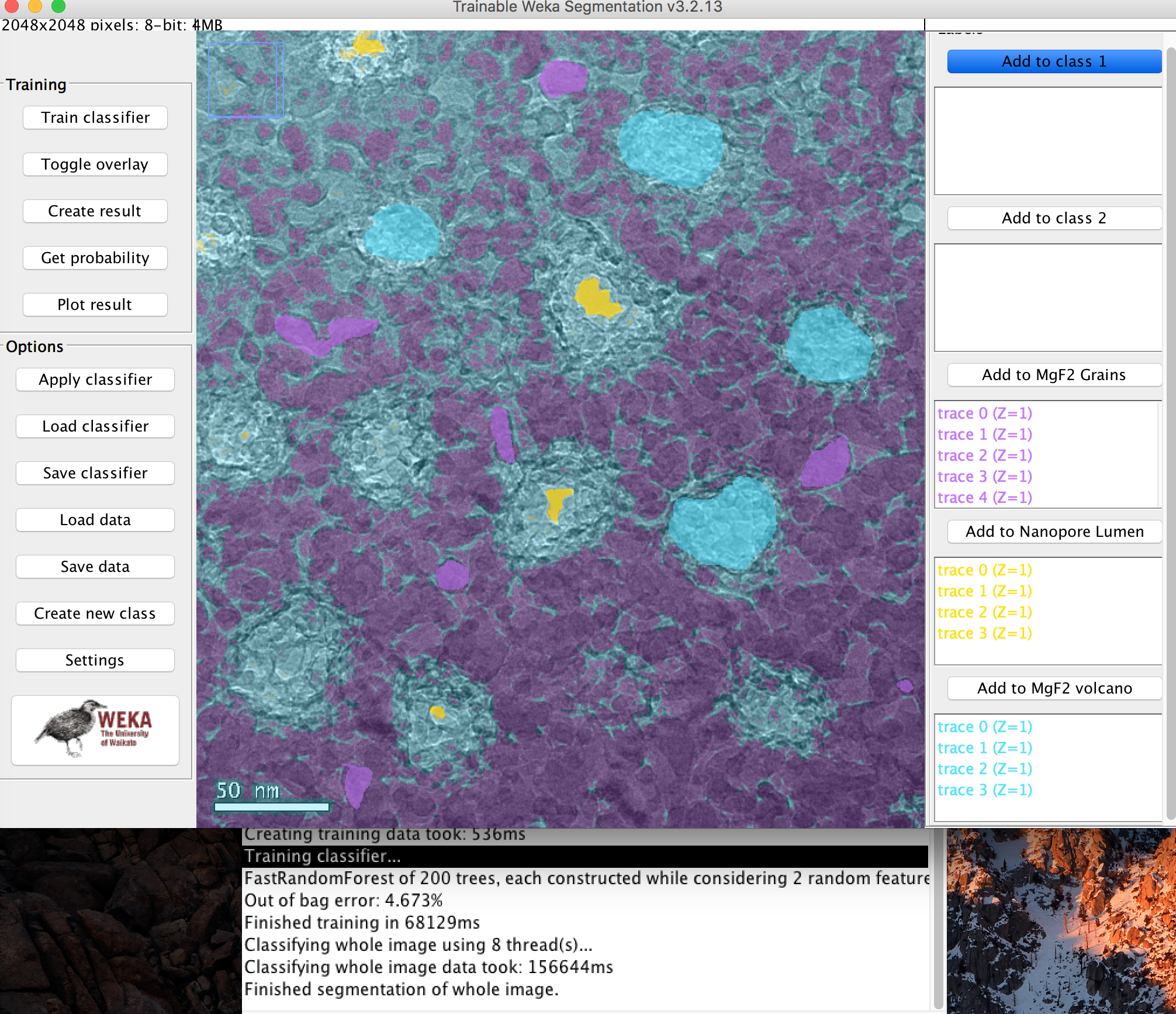
Example Probability Maps
Below are the probability Maps associated with the classifier. You can see definition in the grain that simple thresholding could not achieve.
These maps can be used in conjunction with pore processing software to generate histograms of interesting features.
