BMES 18 – Exosomes: separation and sensing
Hello everyone,
Aslan and I attended talks regarding EVs enrichment which can be divided into two broad categories depending on their ability to recover and reuse the EVS or not. There is a recent review called “New technologies for analysis of extracellular vesicles” published this year on Chemical Reviews. Their breakdown of the different methods is a good introduction for this post.
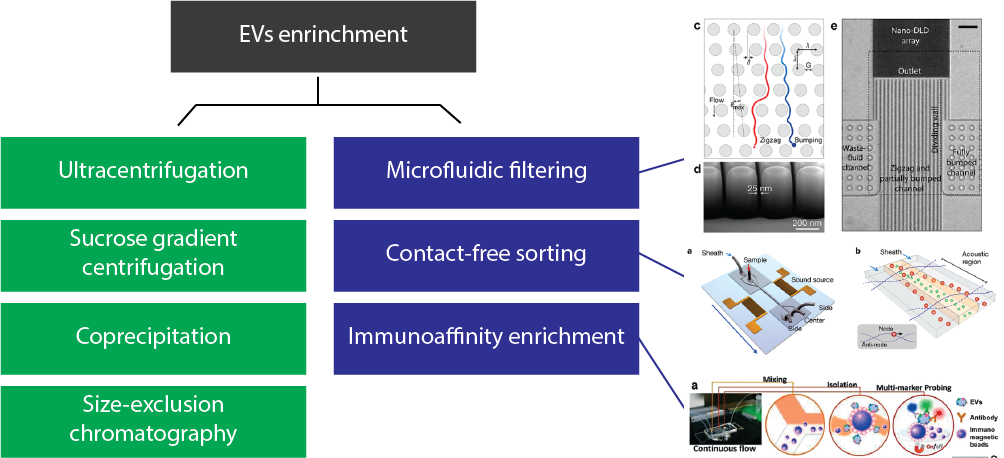
The table below summarizes the traditional methods (green in the schematic above) and their advantages/disadvantages. It seems they exist within a quality vs time trade-off.
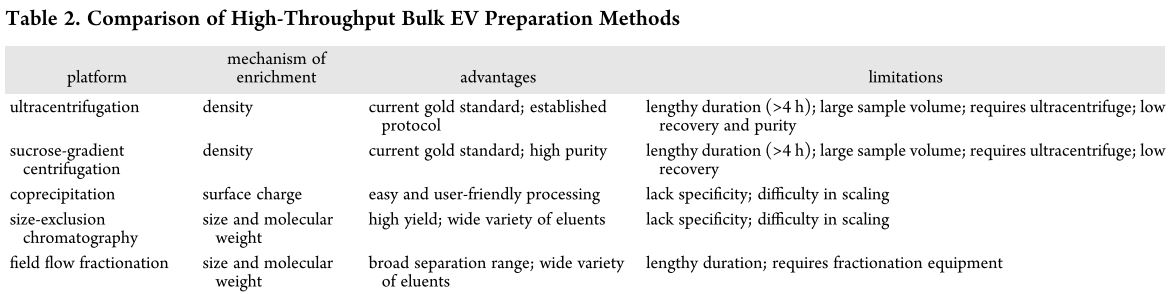
Alternative methods presented at BMES 18
Contact-free sorting
1. High-Throughput Acoustic Separation of Circulating Tumor Cells – Abstract
Mengxi Wu,1,2 Po-Hsun Huang,2 Chuyi Chen,2 and Tony Hunag2
-
- Pennsylvania State University, State College, PA,
- Duke University, Durham, NC
Here you can find the lab website Website. So the speaker was the first author of my qualification exam paper, so i really wanted to go to his talk and ask couple of question about the exosome separation using this technique which they have actually published a paper on it. My qualifying exam paper was proof of concept paper using fluorescent particles but last year they published another paper showing separation of exosomes. The talk was about acoustic separation of circulating tumor cells, and they still needed to prefilter their sample, and they said the are not able to run the whole sample. And when i talked to the speaker, they said that purification of exosomes and proteins is still tricky for them. It seems that this technique is good for separation of 2 distinguished size of particles, but not a mixture of different size of particles. Or at least, they are not able to do that in a single step.
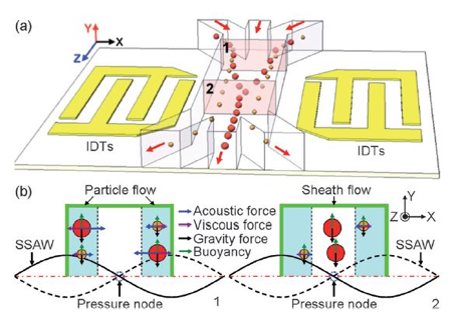
2. Liquid Biopsy for Early Cancer Detection – Abstract
Hyun Ji Kim,1 Fehmi Civitci,1 Josiah Wagner,1 Pavana Anur,1 Matthew Rames,1 Xiaolin Nan,1 Terry Morgan,1 Dennis Duran,1 and Thuy Ngo1
- OHSU, Portland, OR
This was a poster presented by Hyun Ji Kim, graduate student. They do exosome separation from whole blood (ultracentrifugation based) then they label the small exosomes with CD36 and CD81 to identify exosomes. They seem to do flow cytometry on a regular basis as quality control.
She explained their project includes 3 different labs all working on cancer detection from liquid biopsy. They are situated at CEDAR (Cancer Early Detection Advanced Research Center) from Oregon Health and Science University.
She also pointed me to this work by one the PIs in the project, Michael J. Heller:
This is a dielectrophoresis chip to separate small EVs from whole blood, then the EVs are labeled with exosome specific antibodies for cancer detection. They evaluated blood from 20 patients with pancreatic ductal adenocarcinoma and 11 healthy subjects. Their chip had 99% sensitivity and 82% specificity.
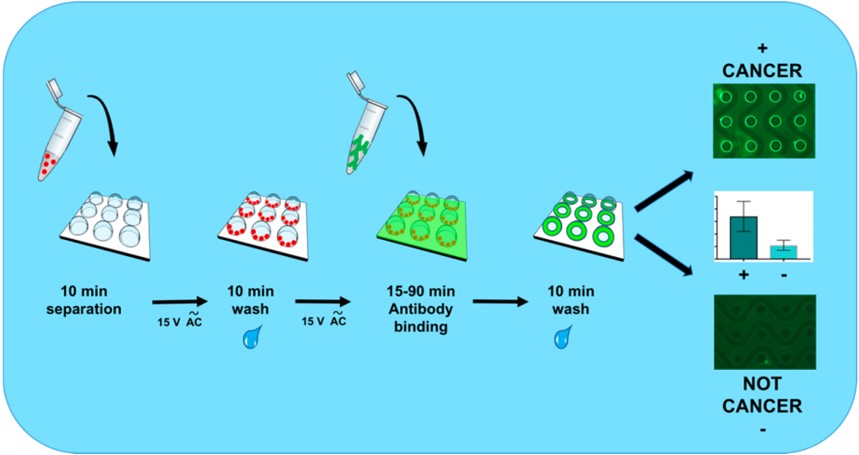
Immunoaffinity enrichment
1. Lipid-Nanoprobe Functionalized Silica NanoStructured Microdevice for Exosome Enrichment – Abstract
Mackenzie Maurer,1,2 Yuan Wan,1,2 Hong-Zhang He,1,2 Yi-Qiu Xia,1,2 Si-Jie Hao,1,2 Wen-Long Zhang,1,2 Nelson Yee,3 and Si-Yang Zheng1,2.4
- The Pennsylvania State University, University Park, PA
- Penn State Materials Research Institute, University Park, PA
- College of Medicine, Penn State Hershey Cancer Institute, The Pennsylvania State University, Hershey, PA
- The Huck Institute of Life Sciences, The Pennsylvania State University, University Park, PA
Project PI: Si-Yang Zheng here is his website, their research focuses on nano/micromaterials for liquid biopsy, EVs, diagnostics, implantable devices, and lab-on-chip. Mackenzie Maurer (gradstudent) presented the work.
A silica substrate which they patterned (by dewetting) to have a random array of pillars. They claim to have control on depth, diameter, and density; however, based on their method that control cannot be precise or broad. The substrate is then functionalized with the lipid-nanprobe (cholesterol-PEG1000) and allowed to interact with a sample containing exosomes (MDA-MB-231 and Panc-1 cell lines spiked-in plasma). The lipid-nanoprobe captures the exosomes and they characterize DNA content after lysis. They haven’t published this work, but they have previously tried a similar approach using Si nanopillars.
The increased surface area increases interaction and the nanopattern can be made such that exosomes are trapped within the pillars which also increases capture and labeling.
Question form the audience: Could you use a cleavable probe to be able to recover the exosomes?
Answer: Good idea, that would be interesting
They recently published a paper called “Rapid magnetic isolation of extracellular vesicles via lipid-based nanoprobes” on Nature Biomedical Engineering (2018) where the attach the probe to magnetic nanoparticles and use a magnet to recover the exosomes for downstream analyses.
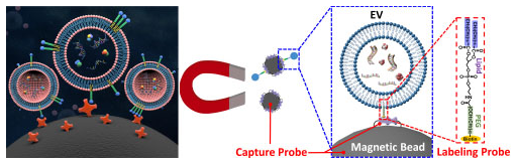
2a. Exosomal microRNA Detection via an Immuno-Biochip for Early Stage Nonsmall Cell Lung Cancer Post- Surgical Surveillance Abstract
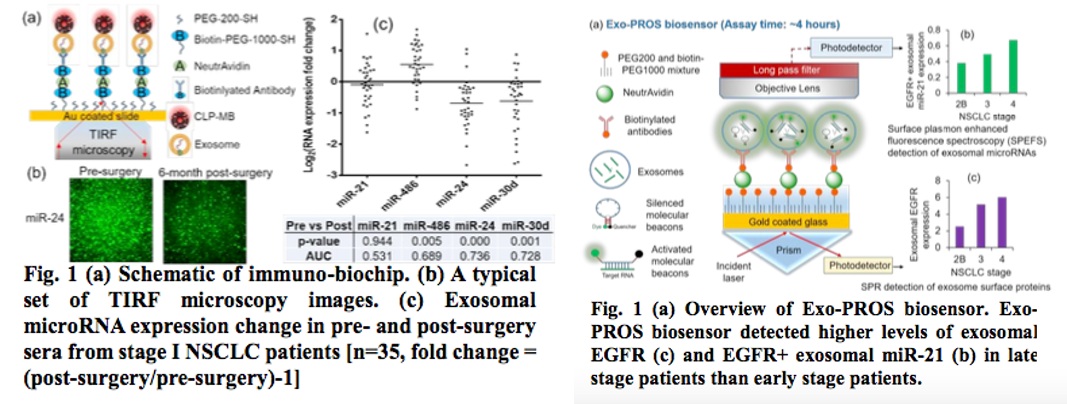
2b. Exosomal Protein MicroRNA OneStop Biosensor: a New Liquid Biopsy Device for Cancer Screening and Early Detection Abstract
Yunchen Yang,1 Eric Kannisto,2 Mary E. Reid,3 Guan Yu4, Santosh Patnaik,2 Yun Wu1
-
- Department of Biomedical Engineering, University at Buffalo, The State University of New York, Buffalo, NY;
-
- Department of Thoracic Surgery, Roswell Park Cancer Institute, Buffalo, NY
-
- Department of Medicine, Roswell Park Cancer Institute, Buffalo, NY
- Department of Biostatistics, University at Buffalo, The State University of New York, Buffalo, NY
Project PI; Here you can find her website Lab Website, their research focuses on: (1) in vivo: multifunctional nanoparticles to deliver imaging reagents and therapeutic reagents for disease imaging and therapy; (2) ex vivo: nanoparticles based biosensor for cancer early detection and surveillance; (3) nanotoxicology: the effects of nanoparticles on disease development.
The PI published a paper for the same application in 2013 when she was doing her PhD, which can be found Detection of Extracellular RNAs in Cancer and Viral Infection via Tethered Cationic Lipoplex Nanoparticles Containing Molecular Beacons.
They have developed a sensitive and rapid IVD assay, i.e. immuno-biochip, to detect exosomal microRNAs for lung cancer post-surgical surveillance. Exosomal miR-24, -30d and -486 combined biomarker distinguished pre- and post-surgery samples with high sensitivity and specificity. Also, They have developed a new liquid biopsy device, Exo-PROS biosensor, which enables one-stop detection of exosomal proteins and microRNAs on a single platform. The Exo-PROS biosensor successfully detected exosomal EGFR and EGFR+ exosomal miR-21 in cell culture medium and human sera. It showed much better sensitivity than ELISA and qRT-PCR.
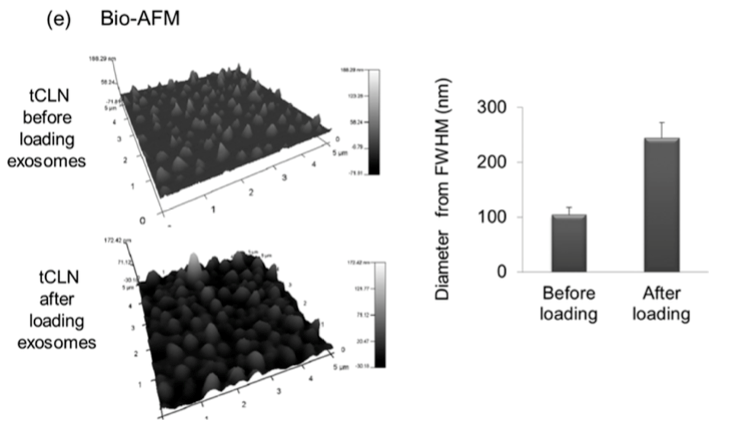
This group could be a potential collaborator, I have been thinking of how to have our membranes in their system. In the talk and the paper, they were talking about fusion of exosomes and lipoplex and i asked them how they characterized this fusion since i have been trying to do this for the PKH paper, and the speaker showed SEM images of before and after and the images looked great, so i talked to him in person and just sent an email to him asking for their protocol. In addition, the paper that PI published in 2013 they have characterized this fusion using Cryo-TEM and BIO-AFM.

Overall highlights
The works presented at the conference focused mainly on “exosome enrichment” and labeling with the common question “can the exosomes be recovered?” Separation protocols that allow for exosome recovery are ideal. Our platform allows for the recovery of EVs in the nanoscale but is not specific to exosomes. Adopting an exosome labeling technique is useful for imaging and analysis but does not achieve the goal of recovering exosomes only
1. Sensing is usually performed on non-porous substrates, with the main drawback of being diffusion dependent. However, a sensor coupled with a membrane is no longer limited by diffusion rates. Since we are able to capture EVs on pores, we could explore the following:
-
-
- Labeling trapped EVs to distinguish exosomes
- Putting affinity probes to selectively capture exosomes (on pores and on the surface), these probes could be cleavable to recover the exosomes
- Ideally, trap EVs on pores, label exosomes/non-exosomes, and selectively recover each fraction
-
2. For the effect of exosomes on wound healing project we could culture cells in DMEM only as well as exosome depleted media and complete media
3. Potential collaborators:
-
- Dr. Yun Wu – University of Buffalo – Onestop biosensor
- Dr. Sabrina Ledlicka – Lehigh University – Isolation of exosomes from cell culture media and NTA
- CEDAR – Oregon