CAD-IA update: GFAP dose-dependency from its non-specific binding to beads
Introduction
As what we’ve learned from the previous “Catch-and-Display Immunoassay” (CAD-IA) trials, we switched to an antibody pair from OriGene with low non-specific binding ability to each other and start to conduct ultracentrifugation for all reactive reagents for the reduction of protein aggregations. However, we found an inconsistency of co-localization percentage of immunocomplexes and beads to GFAP concentrations in the modified CAD-IA protocol. Moreover, we surprisingly found a GFAP dose-dependency when we do not incorporate the biotinylated capture antibodies and came up with a “Catch-and-Display Immunoassay (non-specific binding)” (CAD-IA (ns)), i.e, the assay relies on the non-specific binding of GFAP to the polystyrene nanoparticles (NPs).
Experiments
CAD-IA Several modifications have been made from the previous trials. To reduce non-specific binding of antibodies and proteins to the beads, we used PBS+0.1% BSA as blocking buffer & reaction buffer. The updated protocol is stated below (Figure 1 (a)).
- NPs preparations 2×1010 streptavidin-labeled dragon green polystyrene nanoparticles (d.nm=200, ex / em = 480 / 520) are washed with 100uL PBS (1x) for 3 times to remove pre-added EDTA, anti-microbial, and surfactants.
- “Clean-up” of protein aggregations All reactive reagents including reaction & blocking buffers, proteins, and antibodies are ultracentrifuged by 50,000 rpm through 1 hr before usage.
- Naked NPs captured 8×106 nanoparticles are injected through one port of the uSiM by blocking the other. Remove the tape and wash away non-captured NPs by rushing 100 uL of PBS through the bottom channel and replace solvent in the well with 100 uL fresh PBS.
- Capture antibody binding 100 uL of 2 ug/ml biotinylated mouse anti-GFAP capture antibody solutions were added in both the well and rushed through the bottom channel of the uSiM and react for 2 hrs. The capture ab would be conjugated to NP surfaces based on the sensitive and stable interaction of streptavidin-biotin. Unconjugated capture antibodies are washed away by 3 times with 100uL washing buffer.
- GFAP binding 100 uL of human recombinant GFAP solutions were added in both the well and rushed through the bottom channel of the uSiM and react for 2 hrs. The GFAP will bind to the pre-conjugated capture antibodies. Free GFAP are washed away by 3 times with 100uL washing buffer.
- Detection antibody targeting 100 uL of alexa fluor 647 (ex / em = 652 / 668) labeled mouse anti-GFAP detection antibody solutions were added in both the well and rushed through the bottom channel of the uSiM and reacts for 2 hrs. The detection antibody will target to the pre-caught GFAP and form a fluorescent sandwich immunocomplexes. Free detection antibodies are washed away by 3 times with 100uL washing buffer.
- Imaging the uSiM Switch the laser resource / filters and calculate the co-localization percentage of “red dots” (immunocomplex linked nanoparticles) to “green dots” (all nanoparticles).
CAD-IA (ns) We conducted a control study for non-specific binding ability of proteins & detection antibodies without capture antibody pre-conjugation. The protocol is the same as the one above but simply remove the “capture antibody binding” step (Figure 1(b)).

Figure 1 (a). Current protocol of CAD-IA (with capture antibody conjugation).
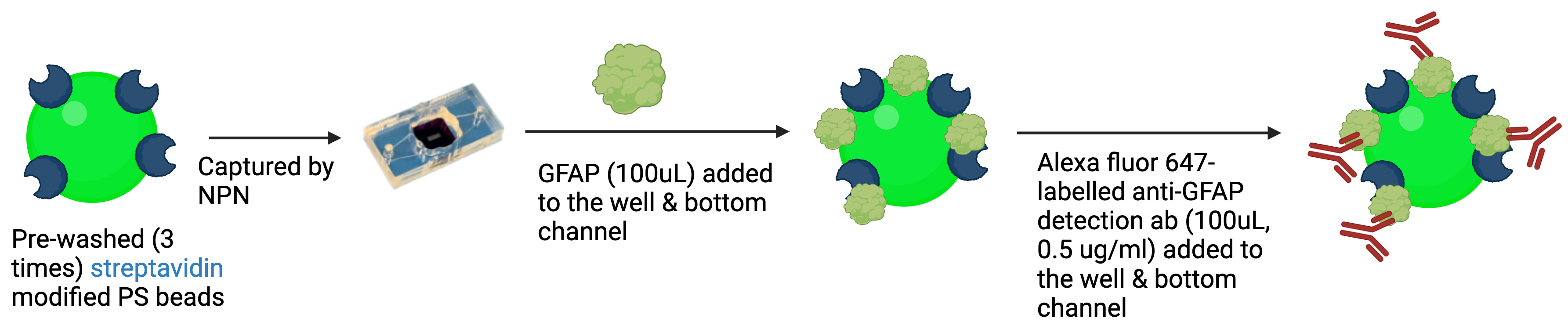
Figure 1 (b). Protocol of CAD-IA (ns) (without capture antibody conjugation)
Results & Discussions
In terms of the CAD-IA assay, capture antibody conjugation efficiency was tested by incubation of a fluorescent anti-mouse secondary igG that specifically bind to the capture antibody (primary mouse igG) conjugated NPs with ~30% co-localization for capture antibody negative and ~ 100% co-localization for capture antibody positive (Figure 2). We conducted a CAD-IA assay with series of GFAP concentrations from (0.07~3.5 ng/uL) and a GFAP negative control (Figure 3). We found that all the GFAP positive groups have higher signals to the GFAP negative groups. However, there was an inconsistency of signal intensities vs GFAP concentrations within the range of 0.07~2.4 ng/uL as they fluctuate within the co-localization percentage of 2~5%. Moreover, we surprisingly found there was a GFAP dose-dependency for CAD-IA (ns) that dose not incorporate capture antibodies in the assay (Figure 4 (a)). The co-localization percentage gradually increase from ~7% (0.07 ng/uL GFAP) and hit the saturation at higher concentrations (>0.7 ng/uL GFAP). This GFAP-dose dependency from non-specific binding of GFAP to the surface of nanoparticles is also found when we further switch the reaction buffer to 10% FBS (Figure 4 (b)) and 100% FBS (Figure 4 (c)), which lost their sensitivity of detection at 0.35 ng/uL and 0.7 ng/uL respectively.
We hypothesized that the inconsistency of GFAP dose-dependency of CAD-IA might result from the poor targeting of GFAP to capture antibody. GFAP will potentially bind to the NPs by specific targeting to the pre-conjugated capture antibodies or non-specific sticking to the surface of the beads. However, the antigen binding domains of capture antibody might be hindered due to its non-specific sticking to the hydrophobic surface of NPs and leads to a poor further targeting of GFAP. As a result, the dominant binding mechanism of GFAP alternatively turn to non-specifically stick to the surface of the beads and compete the binding space with capture antibodies and other blocking reagents to form the observed inconsistency.
For the CAD-IA (ns), the only possible mechanism of GFAP binding is non-specific sticking since there is no capture antibody incorporated. As GFAP concentration increases, more GFAP are involved to compete with other blocking reagents to stick to the hydrophobic surface of NPs. Limit of Detection (LoD) of the assay became poorer as the concentration of blocking reagents increase (0.1% BSA < 10% FBS < 100% FBS) since there is less binding space left for GFAP. The performance of the assay highly depends on the hydrophobicity of the NP surface as we found no intensity signals when we replace the SA-modified NPs with non-functionalized NPs (Figure 5).
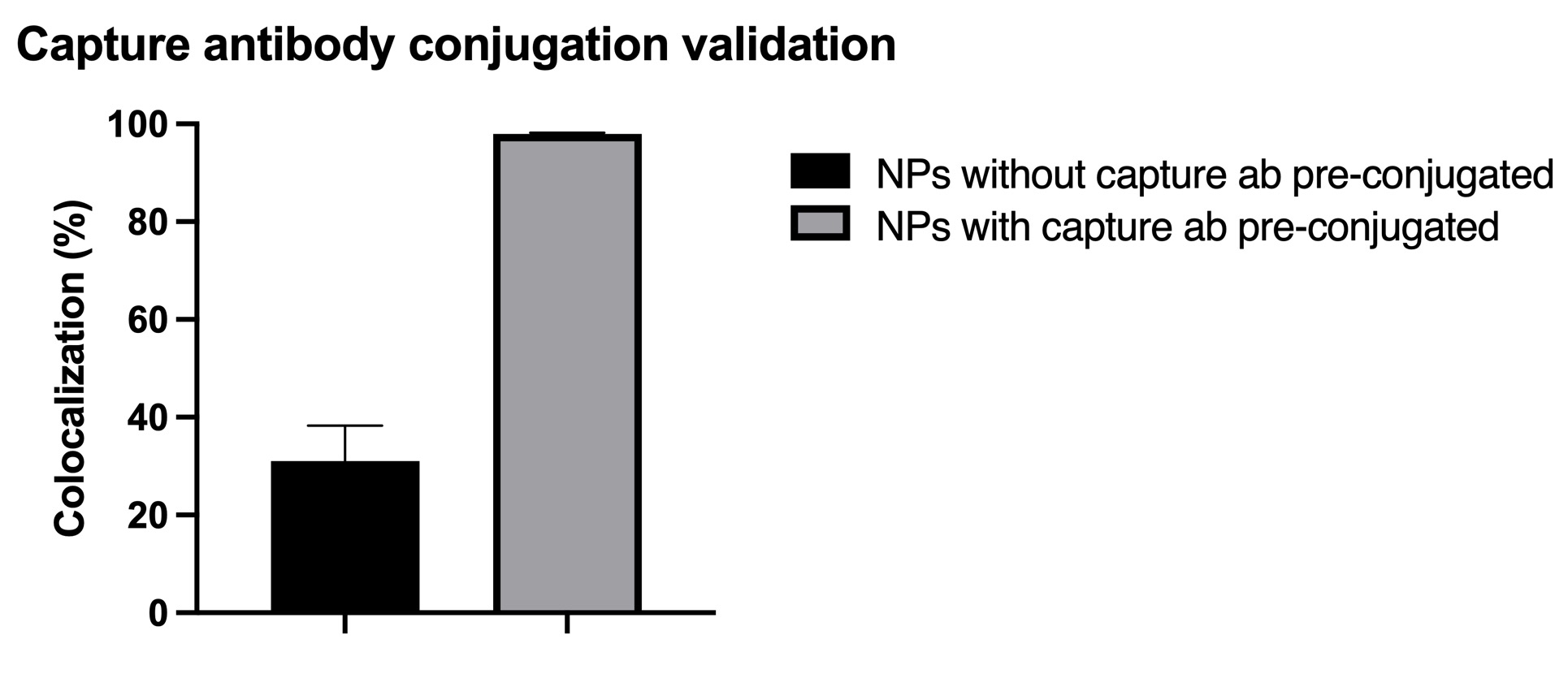
Figure 2. Capture antibody (mouse igG primary antibody) conjugation validation by AF 568 anti-mouse igG secondary antibody. N=6 (6 different regions within the same device).
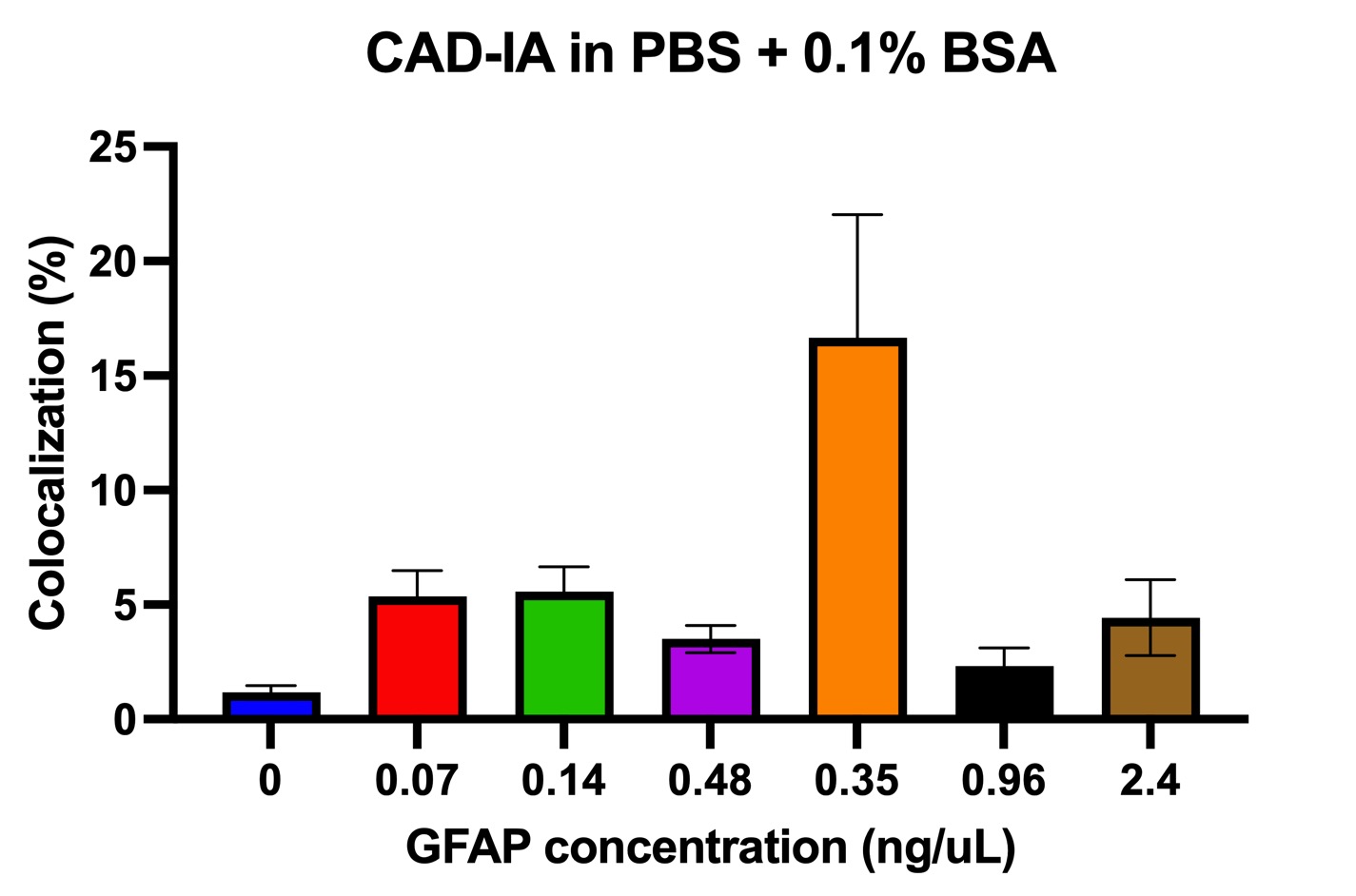
Figure 3. CAD-IA for GFAP detection. N=6 (6 different regions within the same device).
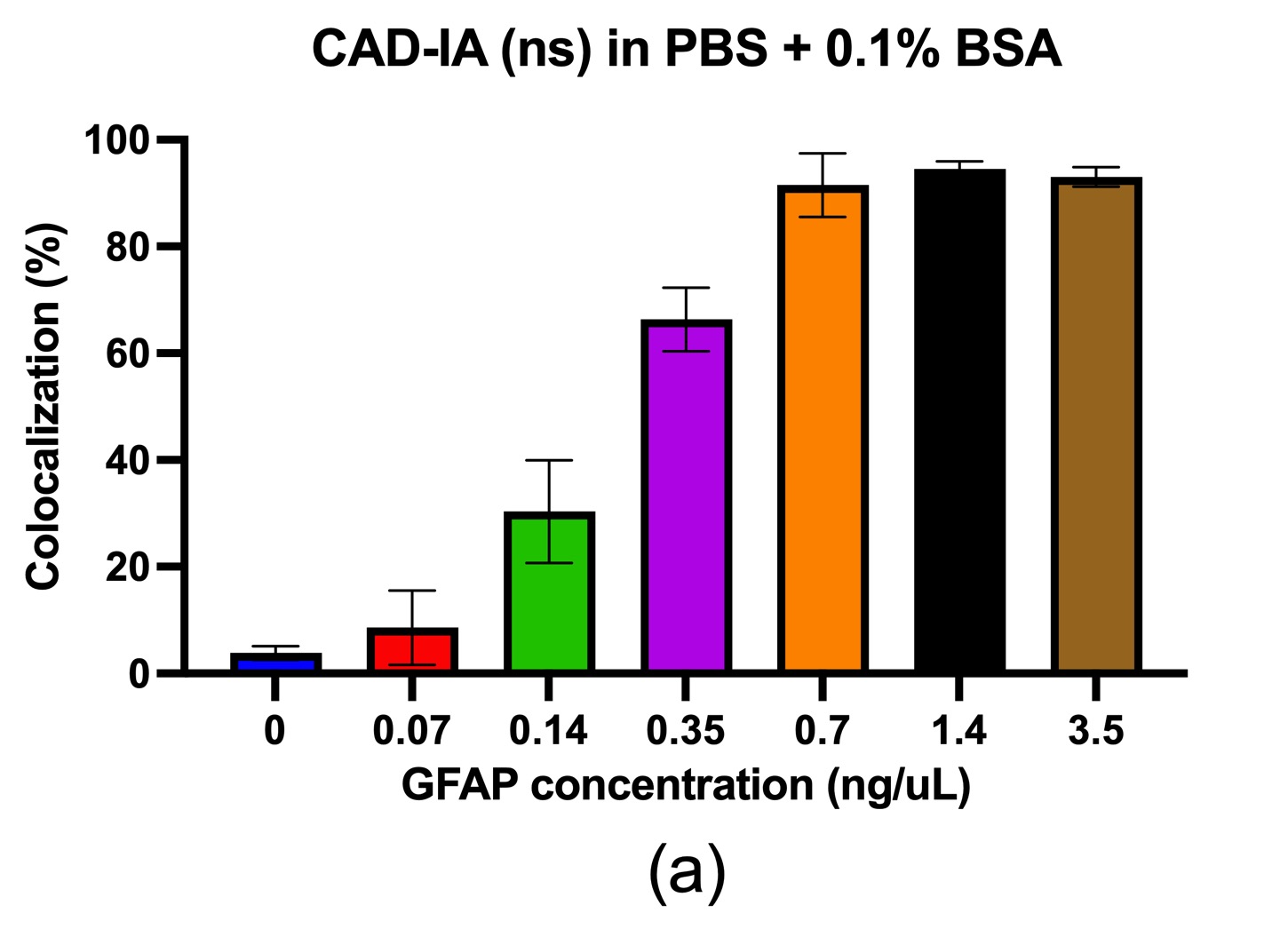
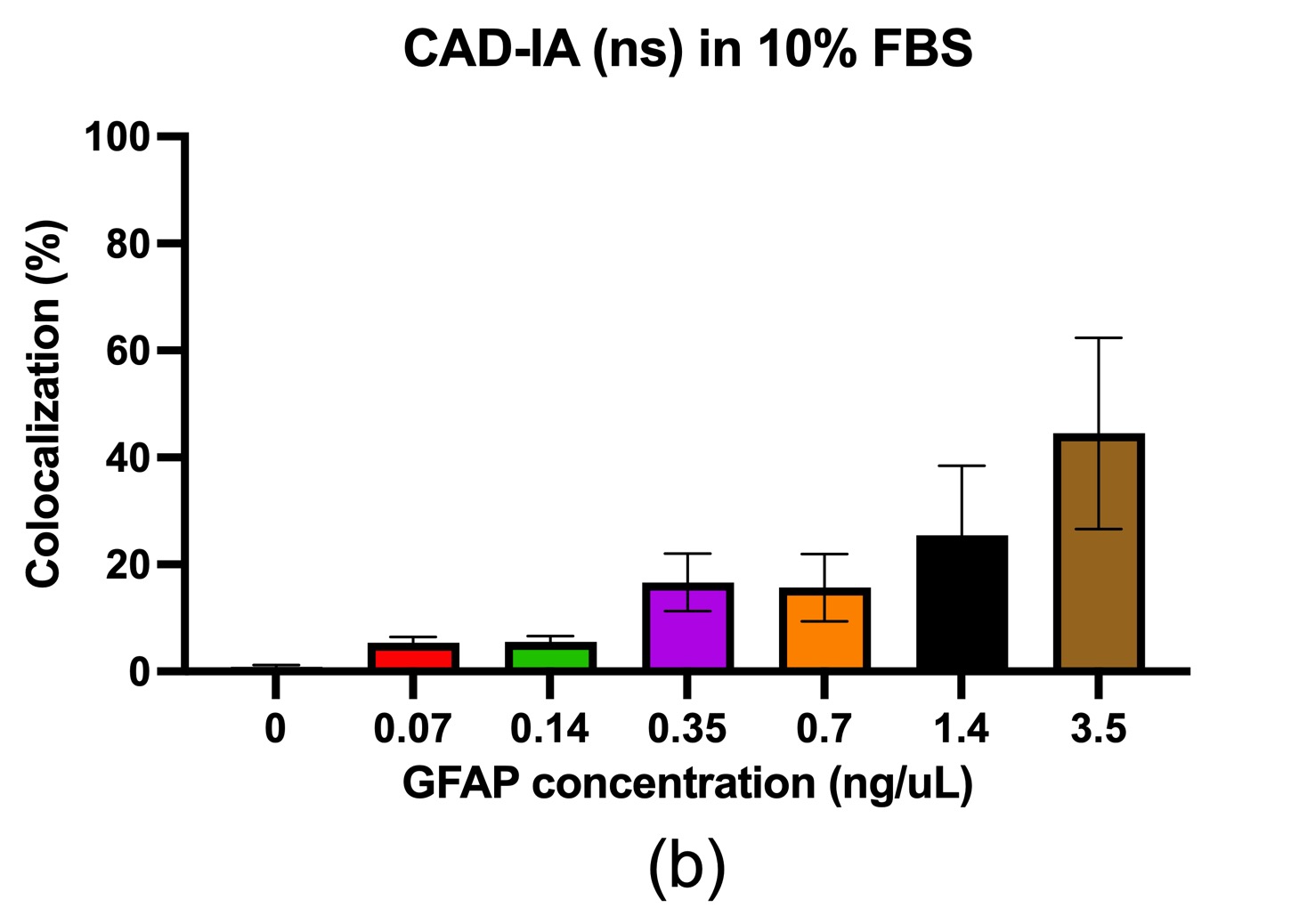
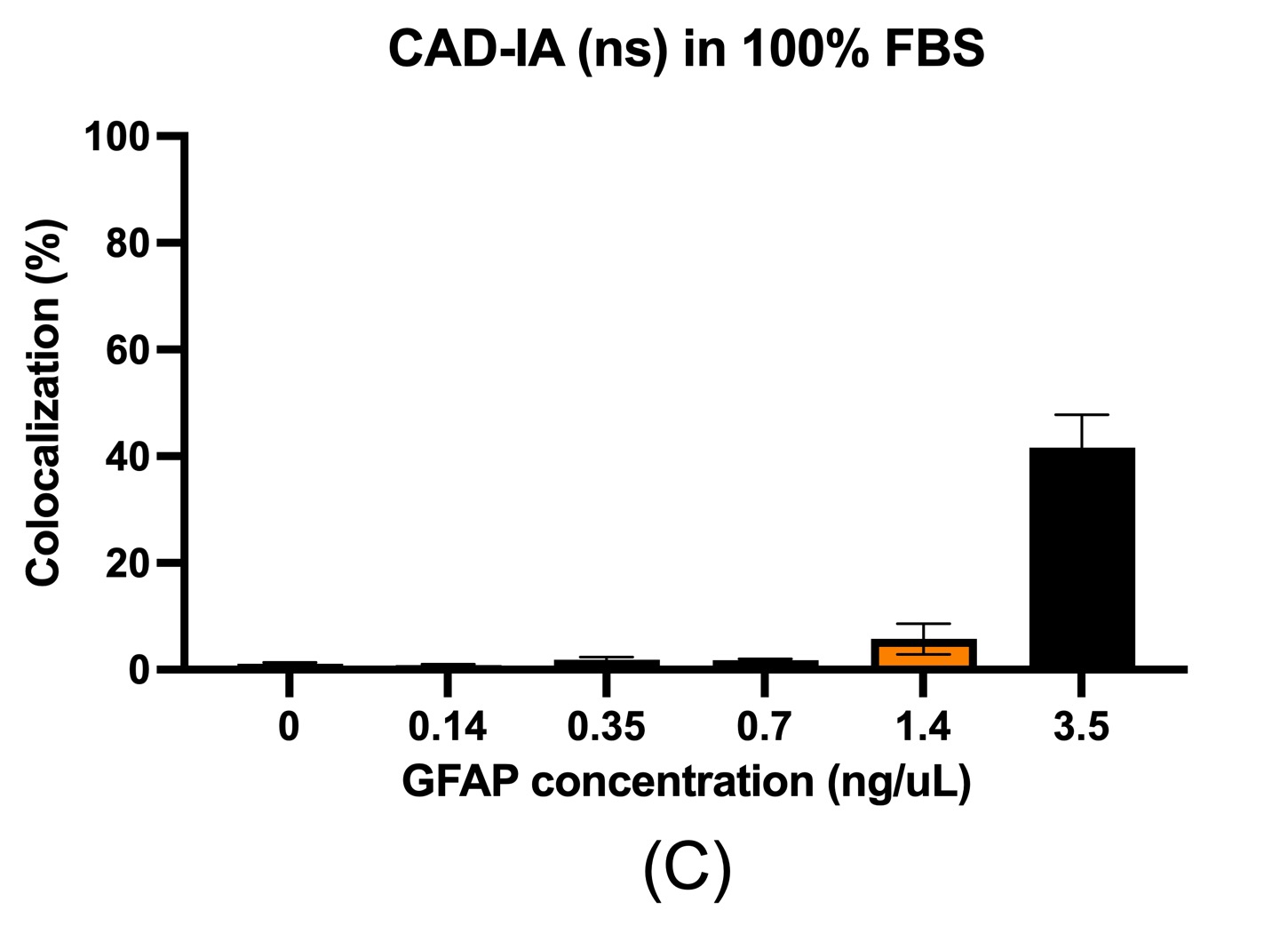
Figure 4. CAD-IA (ns) for GFAP detection. (a) PBS+0.1% BSA as reaction buffer, (b) 10% FBS as reaction buffer, (c) 100% FBS as reaction buffer. N=6 (6 different regions within the same device).
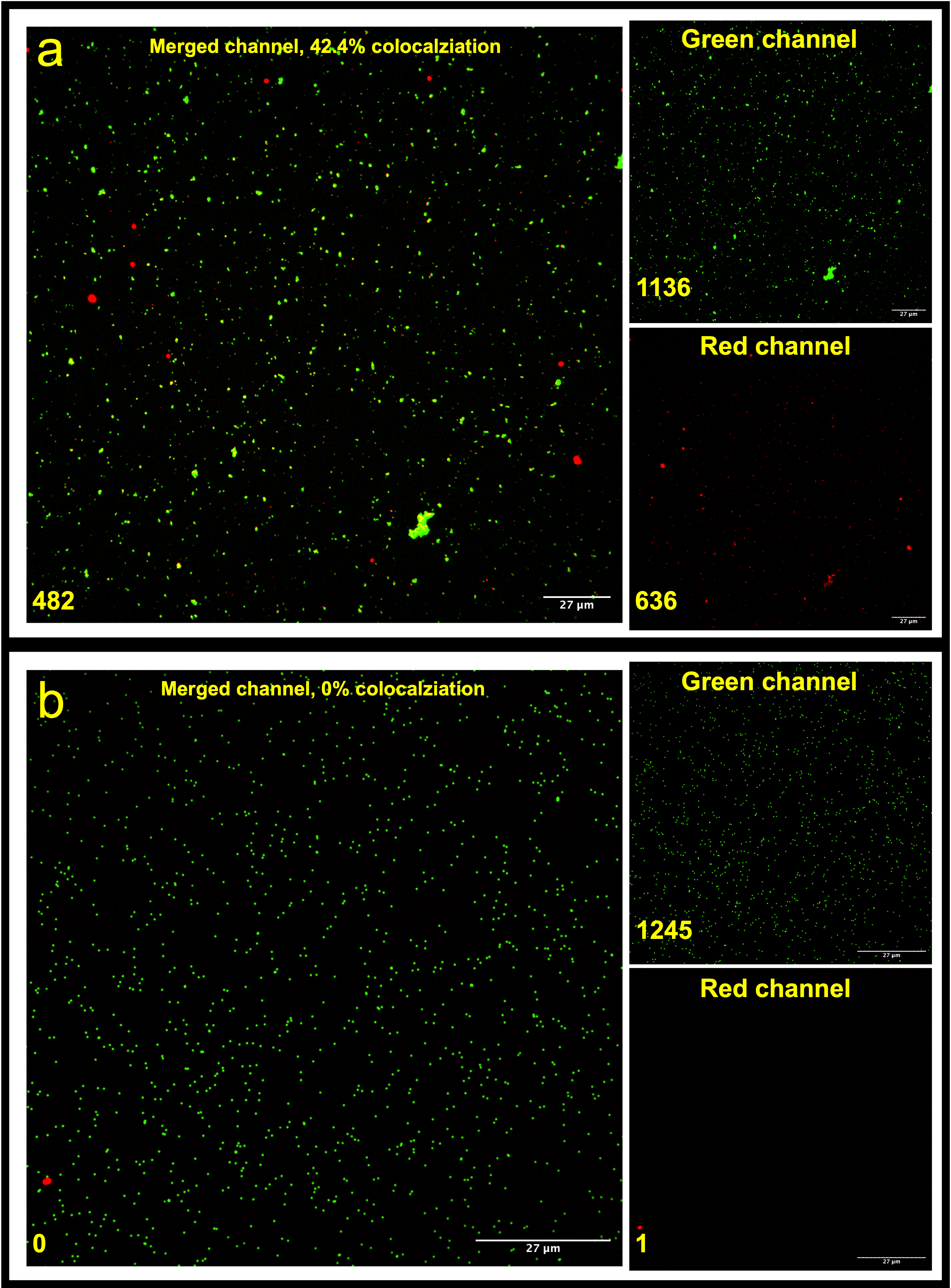
Figure 5. CAD-IA (ns) for 3.5 ng/uL GFAP detection using (a) SA-NPs, (b) naked NPs. N=6 (6 different regions within the same device).
Conclusions & Future directions
The current CAD-IA protocol has been proved with great capture antibody conjugation efficiency and digitally characterize individual fluorescent immunocomplexes but have inconsistent dose-dependency of antigens. We will start to look for other materials with greater non-specific binding resistance (e.g. PEGylated NPs) and better GFAP binding accessibility. On the other hand, the concentration of antibodies, brightness of fluorochromes, incubation time, etc. will be investigated for the optimum reaction conditions.
The CAD-IA (ns) counting on non-specific binding of GFAP has been proved with great GFAP dose-dependency. However, the sensitivity cut-off of the assay (ng/uL) is still 3 magnitudes away from the clinical TBI testable standards (ng/mL)1. We will strive to adjust the reaction conditions to have greater LoD of the assay.
Reference
- Vos, P. E. et al. GFAP and S100B are biomarkers of traumatic brain injury: an observational cohort study. Neurology 75, 1786–1793 (2010).
