14 Days of Cell Growth on PET and pnc-Si transwells
I continued to run some of the samples that I described in this post last week. These are P8 bEnd3 cells, seeded at 100000 cells/cm2 (just below confluence) and grown for 14 DAYS. The pnc-Si was SC126 that was RTP’ed. I measured the TEER of these samples for 2 weeks and then stained the cells on day 14 with Live(green)/Dead(red). About 25% of the original pnc-Si samples remained intact (both membrane slits) for the entire 14 day experiment! (My poor handling broke a few of them, a few of them broke on their own). This is the longest cell culture experiment done with pnc-Si so far.
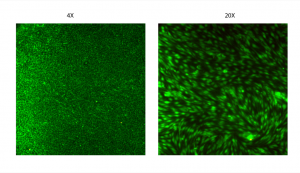
After 1 days on the bottom of PET transwells, there is a beautiful endothelial monolayer (except for the edges, remember) with barely any dead cells (4X and 20X images below):
On pnc-Si, at 4X, there is also a beautiful monolayer, with no edge effects are few dead cells:
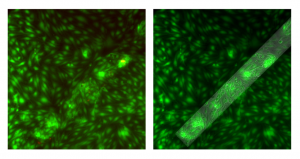
There seems to be an arrangement of cells around the membrane windows – here’s a closer look (20X):
Left is Live/Dead overlay, right is Live/Phase Overlay to outline the membrane window. It looks like some of the cells are aligning at the interface of supported/free-standing membrane. Also, there are those pesky clumps of cells on the membrane but there aren’t as many dead cells associated with those clumps. It seems like the cell clumps are a bit smaller than in my previous post.
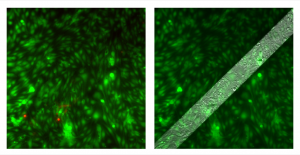
On a different pnc-Si sample (left Live/Dead overlay, right Live/Phase overlay):
There are maybe 2 (maybe 1?) cell clumps over the membrane and the rest of the cells constitute a continuous monolayer. I’m not sure why, in the same experiment, clumping was such an issue on some samples but not others. Was this sample an anamoly?
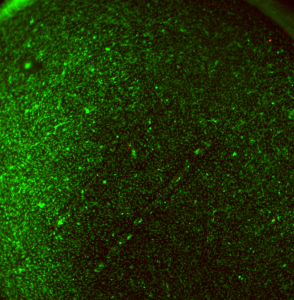
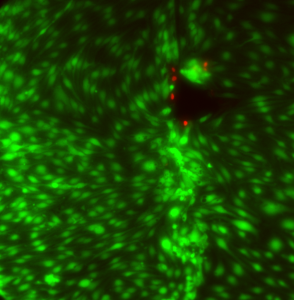
Finally, I got this cool image of cells at a broken membrane area (I broke this membrane during the staining protocol). The cells near the membrane tear are dead. Lots of clumping on this one away from the tear:
This comment is not meant for Barrett’s most recent post only or even at all. I propose that all posts start with a short summary of the results/lessons learned/whatever else. This would be a helpful guide for most readers. In the past I have found that a good number of posts did not have a clear conclusion/lesson learned. Think of what I ask as a short abstract before the main paper. Just a thought but when I am in the middle of a major proposal or review paper or some other task, having such a summary of the post would help a lot! And I think it would help the one who posted it focus on the meaning of the post.
Is it possible to do any confocal mapping of these samples? It seems like some 3-D info would be helpful to understand this clumping issue.
When I look at all these images, it seems to me like the membranes are “prime real estate” for the cells, and they seem to be fighting each other to be there, creating clumps and piles. Is it possible to seed at very low densities where you would not expect a monolayer to grow over the planer surface? If a monolayer were to grow only on the membrane, it seems like it would be useful information.
How about conducting an experiment in which cells are seeded on the supporting membrane but prevented from adhering to the free standing membrane? Over time, cells should grow and migrate towards the free standing membrane? This migration could be compared to a blank chip.
Chris – I’ve been thinking about confocal – especially in light of my recent results with astrocyte multi-layers. I’m not sure where to find one. Next time I split endothelial cells, I am planning on seeding at a lower density.
Anant – I’m planning on seeding at a lower density – how do you think you could prevent adhesion to the free-standing membrane without affecting supported membrane adhesion?
This could be tried by using a small cloning ring over the membrane slits and plating cells around it on an inverted transwell, and removing the cloning ring after the cells have settled or adhered. I am not sure if this will definitely work.