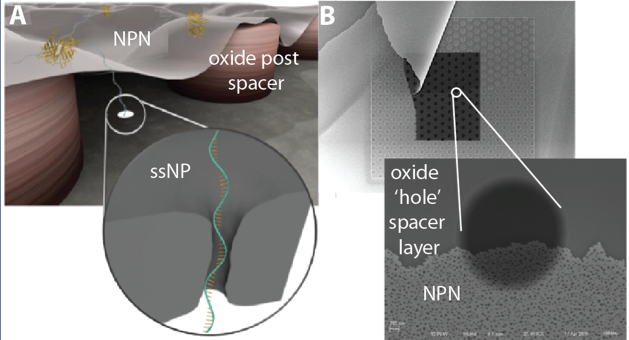
Solid- State Nanopores Integrated with Nanoporous Membranes for Enhanced Single-Molecule Counting of Low-Abundance Biomarkers
We are putting the final touches on a submission to NIH based on our work with Vincent Tabard-Cossa in Ottawa. I want to provide everyone with a small overview.
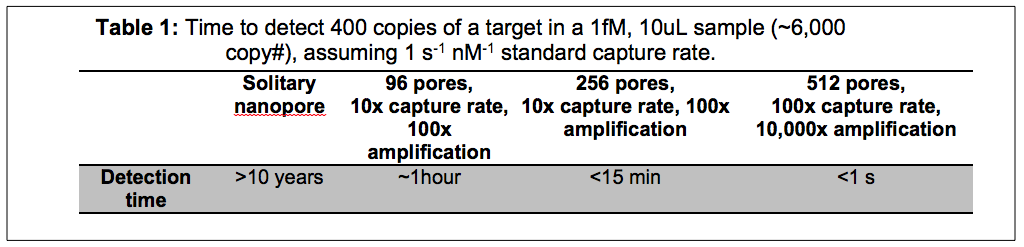
The basic premise is that solid nanopores (ssNPs) are the right tool for detecting low abundance biomarkers for personalized medicine and point-of-care diagnostics. This premise follows from the exquisite sensitivity and processing speed of ssNPs. Because they process molecules one-at-a-time however, there is an enormous throughput challenge to make this dream a reality. Vincent has developed a fantastic illustrating that multiple strategies will need to be combined to make the dream a reality.
 To make matters more challenging, ssNPs are prone to irreversible clogging. This is one reason why they’ve lost the battle to biological nanopores for DNA sequencing. Vincent’s idea was to create a dual chip filter that protected the ssNP from large things that might clog the membrane.
To make matters more challenging, ssNPs are prone to irreversible clogging. This is one reason why they’ve lost the battle to biological nanopores for DNA sequencing. Vincent’s idea was to create a dual chip filter that protected the ssNP from large things that might clog the membrane.
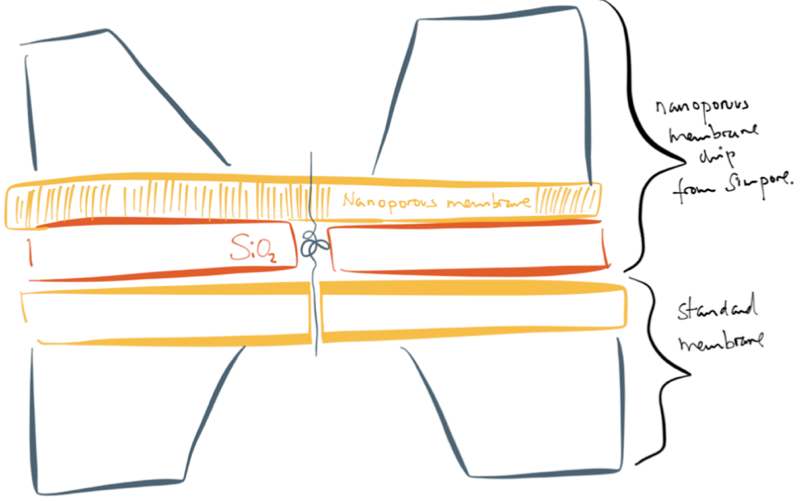 What ended up happening instead of chip bonding was the whole NPN membrane transfer story that you can find archived on the blog in posts from Greg and Tucker.
What ended up happening instead of chip bonding was the whole NPN membrane transfer story that you can find archived on the blog in posts from Greg and Tucker.
The preliminary findings are very encouraging …
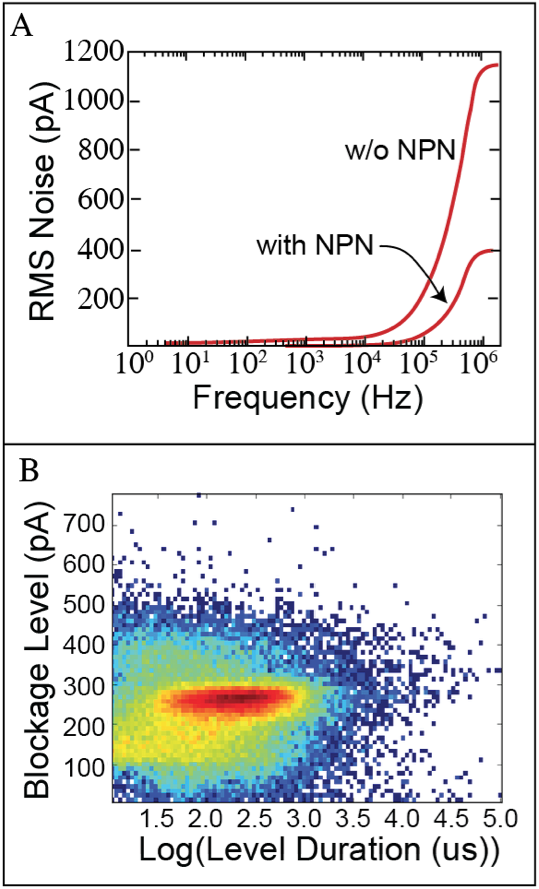 The top figure shows that the presence of NPN reduces the system noise. This allows them to analyze events at 750 kHz, nearly 2x faster than data in the literature. The second data shows the result of the analysis of a record breaking 114,918 DNA events measured continuously from a single pore. This also appears to be more than 2x greater than anything seen in publication – and the system has not been optimized. The grant would do that optimization, integrate the system with microfluidic deliver of sample, and directly test the ability of the NPN/ssNP to help achieve the goal outlined in Vincent’s table. The biomarker amplification strategy we plan to use is shown here …
The top figure shows that the presence of NPN reduces the system noise. This allows them to analyze events at 750 kHz, nearly 2x faster than data in the literature. The second data shows the result of the analysis of a record breaking 114,918 DNA events measured continuously from a single pore. This also appears to be more than 2x greater than anything seen in publication – and the system has not been optimized. The grant would do that optimization, integrate the system with microfluidic deliver of sample, and directly test the ability of the NPN/ssNP to help achieve the goal outlined in Vincent’s table. The biomarker amplification strategy we plan to use is shown here …
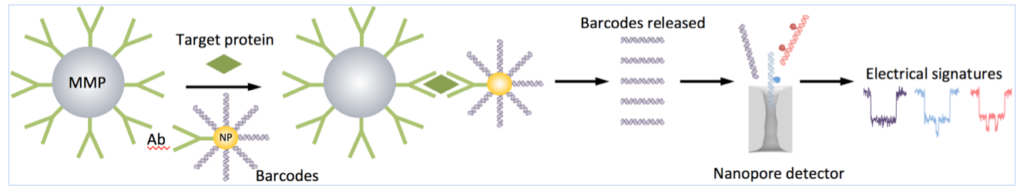 It is not new (Nam, J.M., C.S. Thaxton, and C.A. Mirkin, Nanoparticle-based bio-bar codes for the ultrasensitive detection of proteins. Science, 2003. 301(5641): p. 1884-6.) and may not all that robust frankly, but that is what we are proposing.
It is not new (Nam, J.M., C.S. Thaxton, and C.A. Mirkin, Nanoparticle-based bio-bar codes for the ultrasensitive detection of proteins. Science, 2003. 301(5641): p. 1884-6.) and may not all that robust frankly, but that is what we are proposing.