Super Resolution Imaging of Captured Exosomes
With the installation of the new STED (STimulated Emission Depletion) system in the medical center, I have finally been able to work on imaging individual exosomes (that we know are exosomes because they are labeled for exosomal markers) that have been captured on our membranes. Now, I won’t go fully into the details of how the technology works, but briefly……….. here’s the Wikipedia page. Seriously though, you basically deplete fluorescence in a donut-shaped region and then collect the photons from the center of the region, which allows you to have a resolution down to about 20 nm (people have imaged the individual components of NPCs though, which are only a few nanometers in size). Therefore, this technology would be perfect for trying to image fluorescently labeled individual exosomes.
These samples were tricky to prepare so I had to do some trial and error to get a working sample for imaging. I pre-labeled the exosomes (a Hansa stock) for CD63 in solution and pre-blocked the membrane with goat-IgG. I then ran a capture experiment and labeled with a fluorescent secondary (STAR Red, which is an Abberior dye, the company that makes the STED). These samples were then mounted to a slide and coverslip using ProLong Diamond and covered to avoid photobleaching.
The STED images in two modes: confocal and STED. This allows for the direct comparison of a region with the lower resolution confocal method to STED of the same region. There is a 100x oil immersion objective on the scope and the system has a digital zoom that can image areas probably smaller than single square microns (the smallest that we imaged was a 6 μm x 7 μm area) up to about 75 μm x 75 μm. While running samples, most of the images that I took were high mag (the smaller area) with a 15-25 nm/pixel resolution. These images took only seconds to capture and comparing them side-by-side we can see a stark difference:
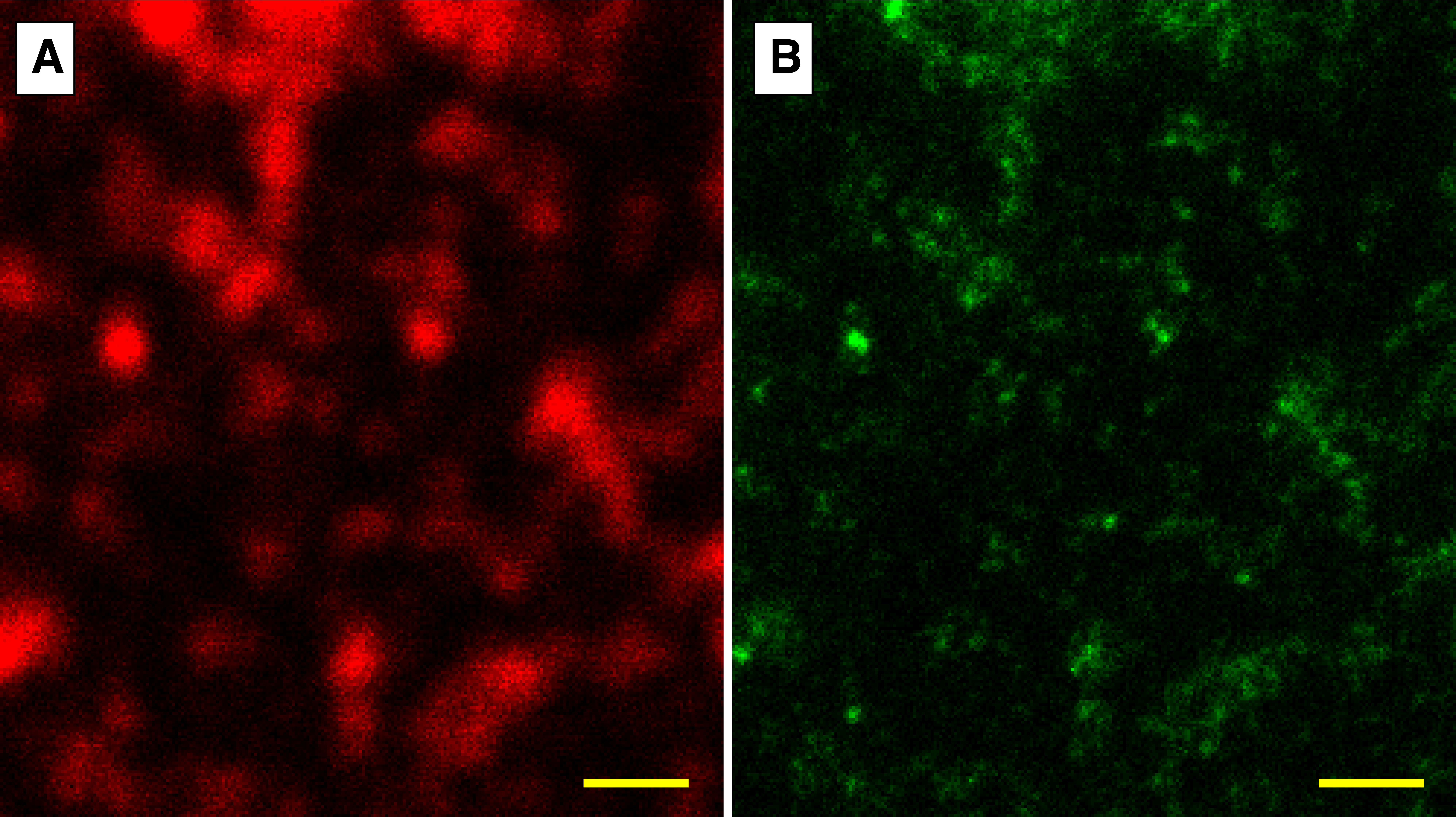
Figure 1: [A] Confocal vs. [B] pseudocolored STED imaging of CD63 labeled exosomes captured on a NPN membrane. Scale bar = 1 μm.
We then took a lower mag image of the membrane at the same resolution (15 nm/pixel) to see a larger area.
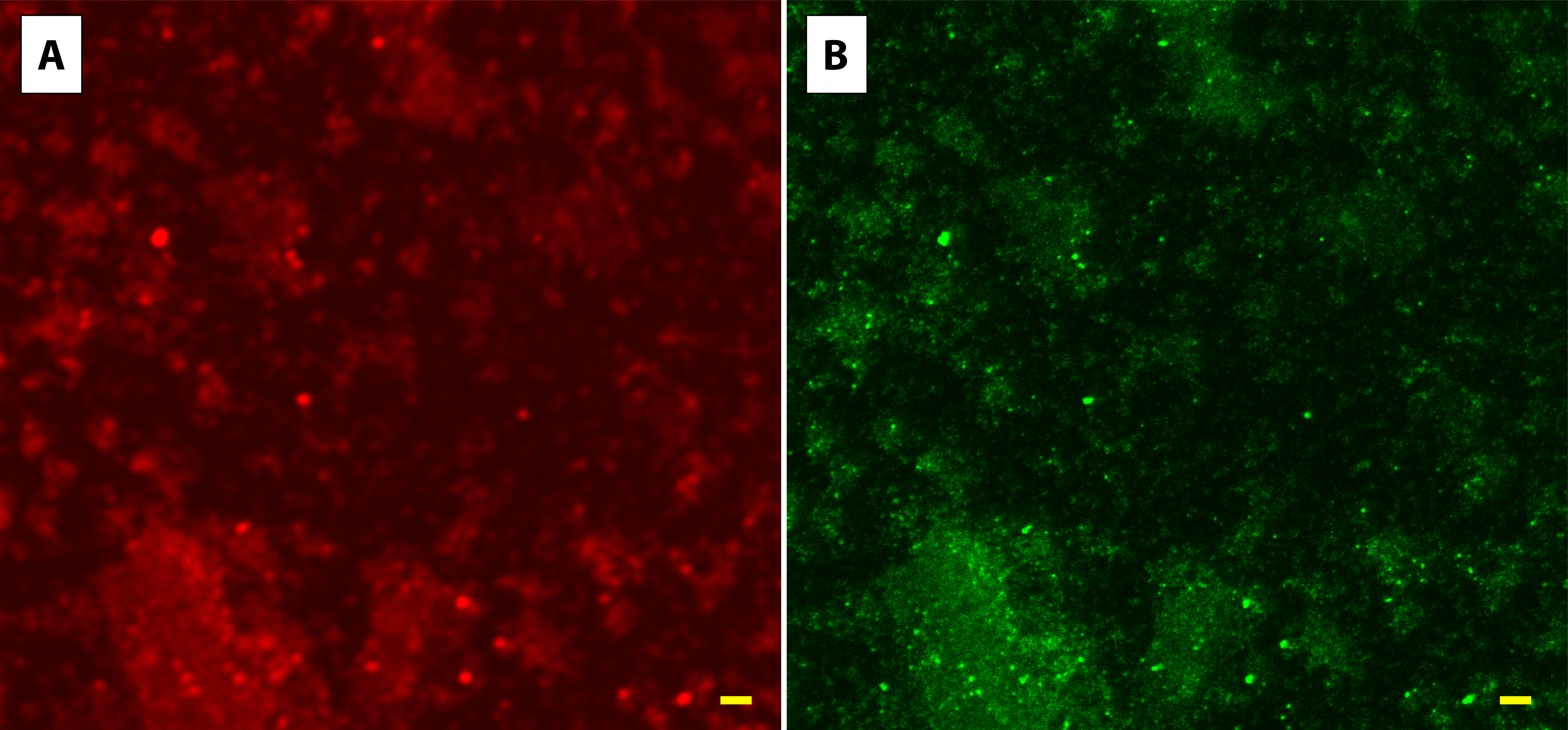
Figure 2: [A] Confocal vs. [B] pseudocolored STED of a 25 μm x 25 μm area with CD63 labeled exosomes. Scale bar = 1 μm.
As we can see, this imaging technique offers superior resolution over stander confocal microscopy, allowing us to resolve what structures are individual exosomes rather than seeing things as more of a fuzzy fluorescent image. There are a couple of issues, namely the fact that we are actually imaging fluorophores that are bound to an exosomal surface protein, so if there is a low copy number of that protein on a specific exosome, then the fluorescence would be low. However, at the higher magnification we should still be able to detect single fluorophores. I did order new sets of dyes (they only gave us a small volume for trying this process out) so hopefully I can label two targets concurrently (MUC2 and CD63) and I also plan on trying a membrane dye to get the exosome only fluorescence (to combat the low copy number possibility) with perhaps BODIPY or the DiO, DiD dyes (at the suggestion of the group in Chicago who have done this kind of thing).