Semantic Segmentation for Neutrophils on HUVEC Monolayers: A Machine Learning Approach
Introduction
Phase-contrast microscopy is widely utilised in live cell imaging, with pixel segmentation protocols in particular finding widespread usage as a means to computationally isolate cells from a tissue-culture background. Computer segmentation allows for the analysis of individual cell properties such as state or spatial location through automated or semi-automated techniques, ultimately attempting to avoid the tedium associated with manual counting. With respect to leukocyte-endothelium interactions, the µSIM physiological systems utilized by our group provides an added layer of complexity to tissue-culture model image analysis due to facilitating neutrophil seeding on a layer of endothelial cells.
Computational segmentation of these images through traditional methods is impractical, if not impossible due to difficulties in accurately separating neutrophils from the endothelial background. Furthermore, traditional computer vision segmentation techniques on grayscale images typically rely on brightness and contrast variations in order to delineate objects from one another. While this is sufficient for neutrophils that are on the luminal side of the µSIM device, any robust probing or transmigration from the neutrophil results in a loss of segmentation capability due to shifts in 8 bit grayscale value that mimic values for the human umbilical vein endothelial cell monolayer (HUVEC) (Figure 1).
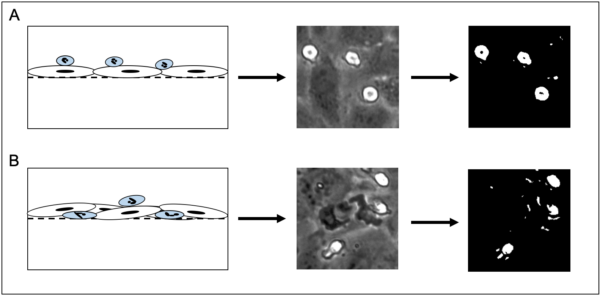
The inability to computationally isolate a neutrophil as it undergoes transmigration effectively cripples our ability to automatically assess neutrophil state (spatially located either luminal/abluminal) in response to chemokine or cytokine stimulation. Furthermore, individual cell tracking is made difficult by an inability to assess the transmigrated state, necessitating manual tracking. While cell tracking can be enhanced through the usage of fluorescent staining, it is important to note that live cell staining still fails to address bulk assessment of neutrophil state.
To this end, computer vision techniques have steadily improved over the last decade. Semantic segmentation via machine learning presents unique advantages with respect to pixel level segmentation of biological imaging samples. This work utilizes a FIJI implementation of the Waikato Environment for Knowledge Analysis (WEKA).
Materials and Methods
FIJI supports the WEKA segmentation plugin that allows for ground truth labeling of images, feature selection, model training, model generation, and subsequent classification through a headless interface via terminal (or command prompt). All of the image processing performed for this work was done with multiple computers. As WEKA does not support graphics processing unit (GPU) acceleration, central processing units (CPU) were exclusively used for computational processing. The main computer used (CPU/RAM/GPU/OS) is as follows:
Ryzen 9 3950x, 64gb DDR4 RAM, Nvidia/EVGA RTX 2070 Super, Windows 10
Beanshell, ImageJ Macro language, and Batch were utilised for scripting, implementation, and automation of the machine learning process via WEKA. Mathematica via Wolframscript was used for data analysis, video correction (described in: ) and plotting. Ground truth labeling was performed in FIJI, with labeled regions saved via “ROI Manager” (Figure 2). Four classes were established in order to differentiate neutrophils and HUVECs, and images were sourced from brightness corrected frames from all experimental conditions (Pos/Neg control, Luminal/Abluminal TNF-alpha, and fMLP). Neutrophils could be in “phase bright” (bright white), “probing” (gray), or “phase dark” (transmigrated, black) while HUVECs were given one class: “endothelial background”. Emphasis was placed on labeling boundaries between classes in ambiguous cases. The four classes were pixel balanced in order to prevent overfitting for one class and multiple features were utilized in order to enhance the model. These include: Gaussian blurs, Sobel filters, Hessian matrices, Difference of Gaussian, Membrane Projections, Mean, Minimum, Maximum, Variance, Entropy, and Neighbors. The mathematical descriptions of these convolutions are described in the ImageJ wiki: (https://imagej.net/Trainable_Weka_Segmentation.html#Training_features_.282D.29).
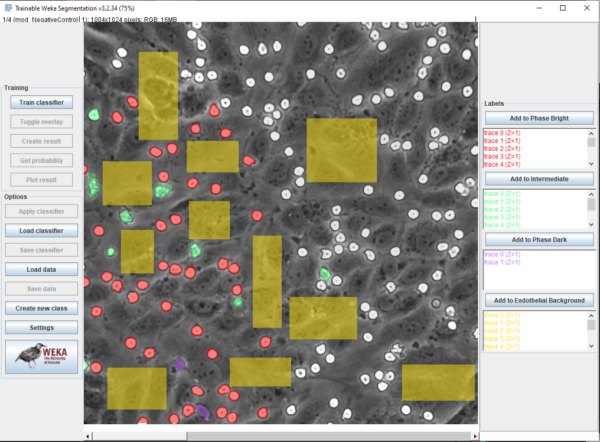
After labeling, models were trained and saved as both “.arff” files (containing ground truth labels/traces) and a “.model” file that was used for video classification. A beanshell script was adapted from the ImageJ wiki in order to process frames of a video separately in order to mitigate memory issues.
Results
After training, the model reported an error of 1.58%, indicating an overall accuracy of 98.42%. This is tested by applying the trained model on the ground truth labeled images, and assessing pixel detection accuracy against manually labeled pixels. The trained model can then be used on microscopy videos, with an example shown below (Condition: Position Control).
Results from a classification procedure include a segmentation map that assigns a pixel one of four colors (correlating to the four classes) based which class has the highest probability of occurring and a probability map of each class (Figure 3).
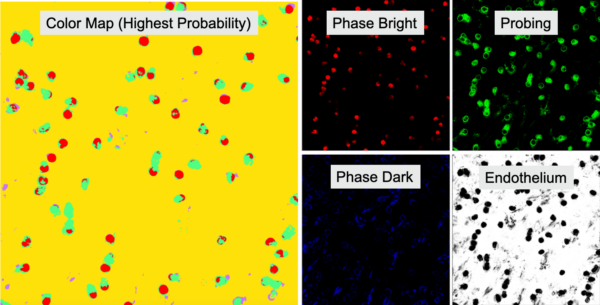
Discussion/Conclusion
Machine learning based segmentation via WEKA allows for robust pixel level classification and computational separation of neutrophils from the HUVEC monolayer. The overall accuracy of this model is high, and this facilitates state analysis and tracking (which will be discussed in another post). There are distinct limitations to note, however. First is the spurious detections that can be seen throughout the video. These are more related to aberrations generated during video recording/correction, and may be fixed soon with a migration to a newer microscope. The second limitation to note is that phase dark segmentation is evident, but not as robust as phase bright/probing. This should not limit our ability to analyze state or track cells, and better performance may be possible with implementation of a deep learning based solution called “Unet”. There will be more on this front later on.