1D Diffusion Model and Nature Results
In this post I will explain how I tried to use the 1D diffusion model to check the results we had in the Nature paper. Unfortunately I didn’t get the same results in the simulation and I present some ideas for the discrepancy.
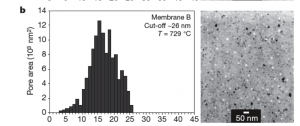
In the Nature paper we made simple small volume diffusion chambers and watched fluorescent molecules and proteins pass through the membrane. We used the following membrane for these experiments:
I reprocessed this image and managed to get close enough results with our pore processor program. The diffusion experiment with this membrane was as follows:
Basically in six minutes, BSA has traveled through the membranes 3 times faster than IgG according to the fluorescence intensity. We know the radius of BSA is 3nm. According to this paper, the hydrodynamic radius of IgG is 5.5nm. If I use the pore processing results files and these two radii, I can find what the concentration would be 50 um from the membrane (the same dimension used in the paper) after 6 minutes.
Results: In the model, BSA is traveling through the membrane 1.31 times faster instead of 3 times faster. Without a membrane (and solely based on free diffusion) BSA would travel through 1.07 times faster. So there is some reduction of IgG diffusion due to the membrane, but not as much.
Suggestions for discrepancy:
1. Geometry. Maybe a 3D system would matter here too, or maybe I’m not checking at the right point for the final comparison.
2. Fluorescence. It’s possible that the fluorescence is not a linear measurement of concentration and this may be altering the comparison.
3. Time. It seems that the 1D model is much faster in general than the 3D model even though I’m pretty sure all the parameters are correct. This may go back again to the geometry argument, but perhaps for a different time point you’d see something closer to the experimental results.
4. Comparison. The experimental difference in hindrance between a membrane and no membrane system would be the best comparison to make, however this is difficult to set up as the membrane works to compartmentalize the system.
5. Pore characteristics. While this needs to be a very big change to affect the results, maybe the pore processing was pore or electrostatics/adsorption affects change the pore sizes.
Comments here would be very useful.

As we showed in the supplemental materials in the nature paper, it’s highly likely that some of the BSA would stick to the membrane, reducing the pore size considerably (a monolayer would close a pore by ~7nm). If you did a subtraction from the diameter of each pore, how much would the pore size need to be reduced to produce the separation characteristics observed in the nature paper?
Do you think the difference between the 1-D and 3-D models has more to do with the increased distance to the membrane, or the fact that so little membrane area exists relative to the area of the fluid? It’s probably difficult to de-couple these effects, but they are distinct. One is just free diffusion to the membrane, and the other is a resistive term created by forcing all the flux through a tiny active area.