Optimization of our isolation Technique using Track Etched Membranes
Hello Everyone,
Following my experiments with Track Etched Membranes:
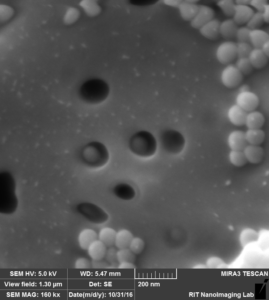
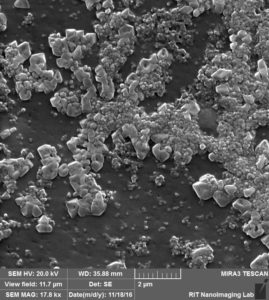
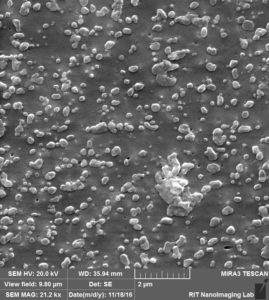
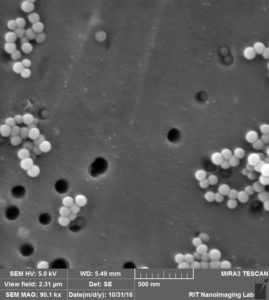
So we came up with this idea to perform SEM imaging. Following are the images that i took for my experiment with 100 nm beads over 100 nm pores after releasing step:
So in this image, we can see that there are a lot of beads that are attached to the surface, at the same time there are some beads that are actually inside the pore, probably clogged.
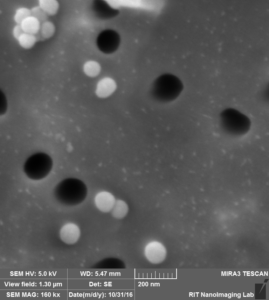
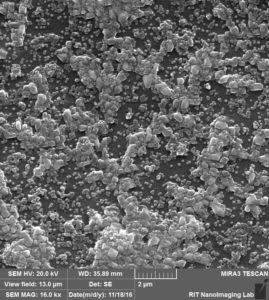
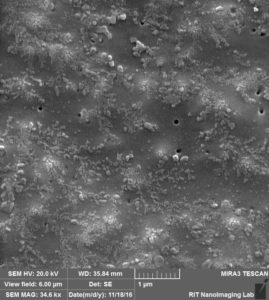
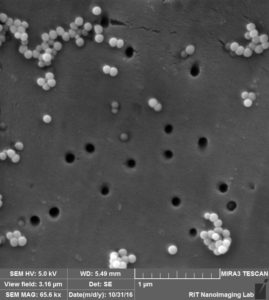
So in the second image: again you can see that beads are attached to the surface, Further more: 1. we can see a lot of aggregation that happens probably during drying s 2. size of the beads and also pores are not exactly 100 nm.
So for surface attachment, i am gonna load the channel by 30 microgram/mL Albumin for 30 minutes to cover the surface of the membrane for decreasing the surface charge.
My plan is to run 4 experiments with 10 nm and 80 nm pore sizes of TE membranes with and without Albumin. (using 100 nm Beads)
1. 80 nm pore without Albumin:
Expectation: capturing of 100 nm beads on the 80 nm pores, and also surface attachment of beads.
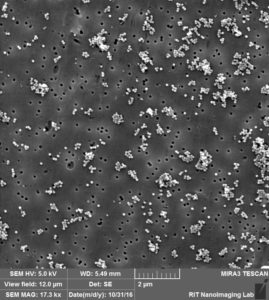
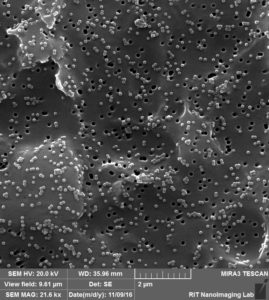
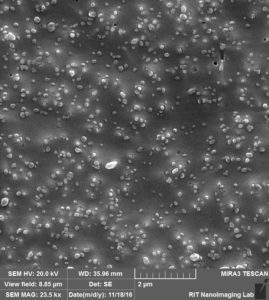
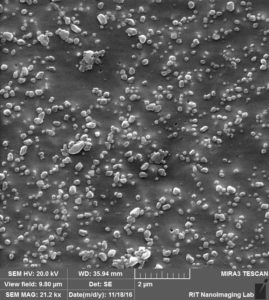
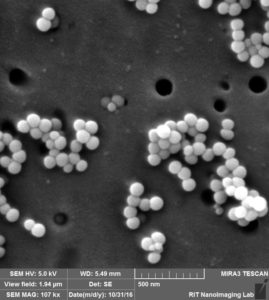
So by these images clearly, 3 different situation can happen:
- Attaching to surface 2. Trapping over pores 3. Clogging the pores.
Like we expected, a lot of beads that are sitting on the membrane because of charge interaction or during the drying. at the same time, we can see beads that are trapped over pores or clogged inside the pores.
2. 80 nm Pore with Albumin:
Expectation: Capturing of 10o nm beads on the 80 nm pore, but no surface attachment due to albumin.
Second Experiment:
Third Experiment:
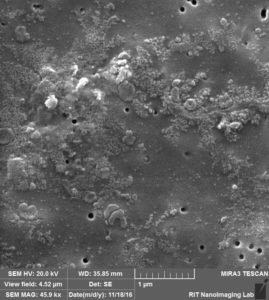
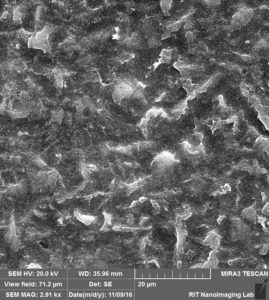
So obviously, Albumin is covering the surface of the membrane and pores in a way that pores are totally buried underneath of the Albumin, another thing to mention is probably salts from PBS that are available over the membrane probably during drying step.
Furthermore, for negative control experiments of this step, i imaged the membrane itself and also the membrane covered by Albumin. The membrane itself is not even close to SEM images that they show on their website, not uniform distribution and a wide range of pore sizes.
The membrane:
The Membrane coated by Albumin:
For the next step:
- I would like to check if the beads are already aggregated in the solution by Zetasizer.
- I would rinse my membrane after my experiments using DI water.
- I would like to check if we can change the zeta potential by PH to control the charge interaction of beads and membrane.
3. 10 nm pore with Albumin: (Negative Control)
Expectation: not to capture any 100 nm bead on the 10 nm pores, and no bead attachment to the surface due to Albumin.
4. 10 nm pore without Albumin:
Expectation: not to capture any 100 nm bead on the 10 nm pores, and all of the signal must be from beads that are attached to the surface.
I will upload my future results and images on this post.