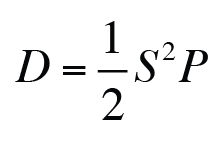BdEC/T24 co-culture: wound healing
Introduction
Motivation: We want to assess if the bladder cancer cell line T24 would influence the baseline characteristics of bladder epithelial cells (BdEC, a.k.a. ATCC® PCS-420-010™) when the two cell types are cultured together. Ultimately, we are interested if the T24 induces BdEC malignancy (turning the normal epithelial cells cancerous). The ultimately test of malignancy would be through invasion assay, but for now, we will look at wound healing first.
Goal: Study how the co-culture of BdEC and T24 cells affect the wound healing of BdEC.
T24 Factoids: According to ATCC, T24 has 19 hour generation time, is tumorigenic in hamster cheek pouch but not in nude mice, and doesn’t fare well in the presence of leukocytes and sera from patients with transitional cell carcinoma [reference]. We grow the T24 in RPMI, supplemented with 10% FBS and 10 U/mL penicillin and 10 μg/mL streptomycin. For short, I will abbreviate this media as RPMI+. Note that the level of antibiotics used is 1:10 the typical dosage. I intendedly do so to match the antibiotics level used for BdEC culture.
BdEC Factoids:
- BdEC takes a day to adhere to the tissue culture plate
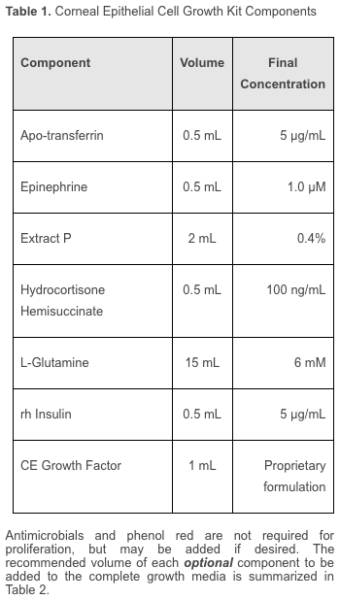
- . BdEC were grown in special media consist of the Prostate Epithelial Cell Basal Medium (PECBM, a.k.a. ATCC® PCS-440-030™) and the Corneal Epithelial Cell Growth Kit (ATCC® PCS-700-040™), supplemented with 10 U/mL penicillin and 10 μg/mL streptomycin. For short, I will abbreviate this media as PECBM+. Note that the level of antibiotics used is 1:10 the typical dosage (in accordance to ATCC recommendation).
The PECBM is serum free, while the corneal epithelial cell growth kit contains the following constituents:
More Factoids: Usually I prefer to grow the cells that need special media in non-special media, since we do not know how the proprietary media constituents will affect the cells under different conditions. However, the BdEC do not fare well in the RPMI+ used for T24 culture (see below).
Hence I have performed all the co-culture studies using the PECBM+ media.
Materials and Methods For details, please see Aslan’s post.
Brief recap here:
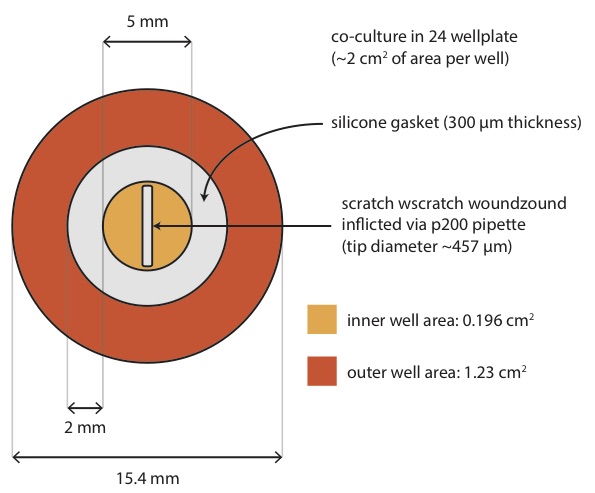
We are looking at about 1:6 ratio (inner:outer) in growth areas, with scratch wound area of about 15% of inner well area.
- I seed the cells at 30% confluency and let grow for 3 days (all cells would double within 24 hr).
- On day 4, I refreshed the media, and let the new media condition for 1 day.
- On day 5, I combined 550 μL of the conditioned media with 500 μL of fresh media (containing 1 μM SiR-DNA).
- I used p200 pipette tip to scratch the wound (1 tip per wound), and wash each scratched wells with 1 mL PBS (with trituration).
- I add 1 mL of the combined media (from step 4) back at each corresponding well.
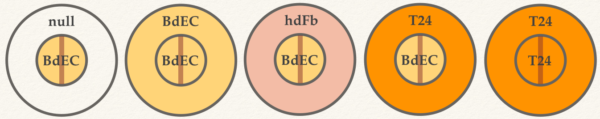
Co-culture combinations (inner/outer):
1. BdEC/null; 2. BdEC/BdEC; 3. BdEC/hdFb; 4. BdEC/T24; T24/T24. NOTES: 1. hdFb = adult dermal fibroblast. 2. We included T24 in inner well too to see how T24 behaves in the PECBM+ media.
With the 10X objective on Leica, 3 fields of view (~1 mm x 1 mm) will completely visualize the whole wound. I image at two non-overlapping fields of view per scratch wound while avoiding the very top and bottom of the scratch.
Results:
Cell density determination
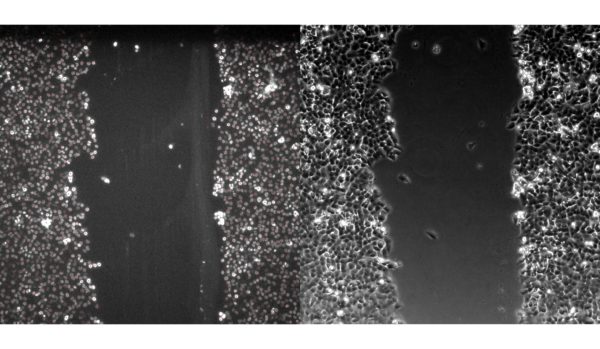
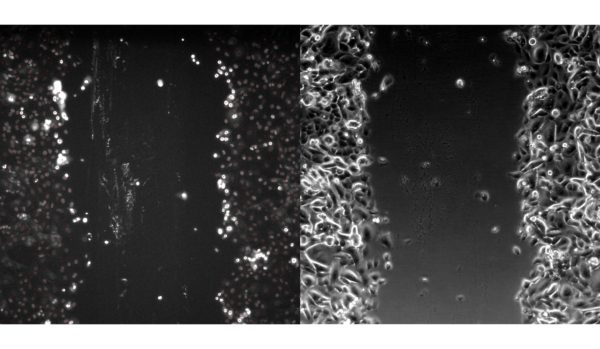
BdEC scratch wound:
T24 scratch wound:
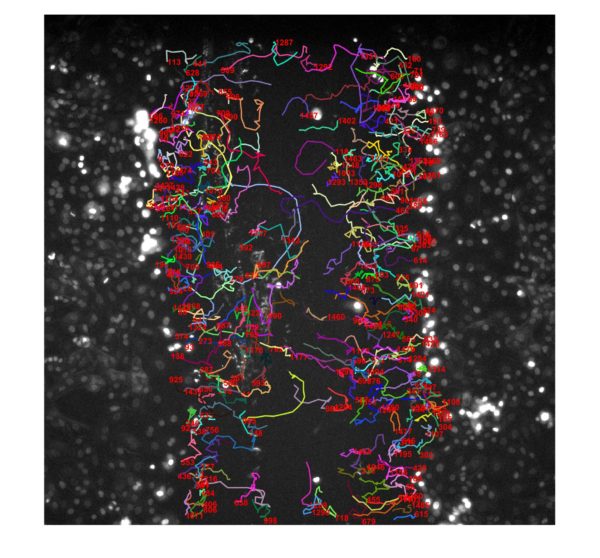
Appreciation of Counting/Tracking Fidelity:
BdEC/T24:
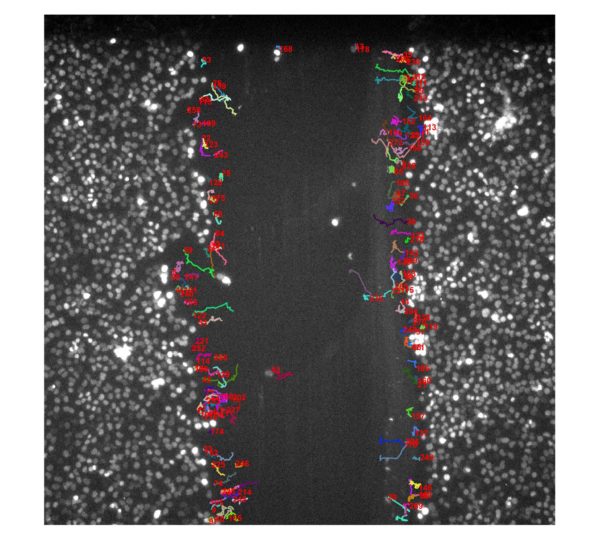
T24/T24:
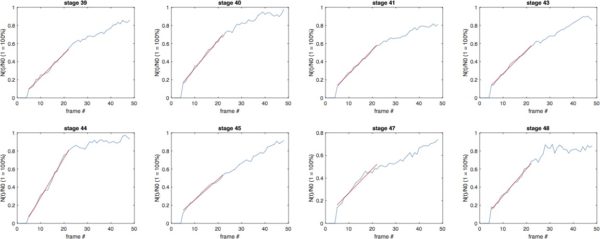
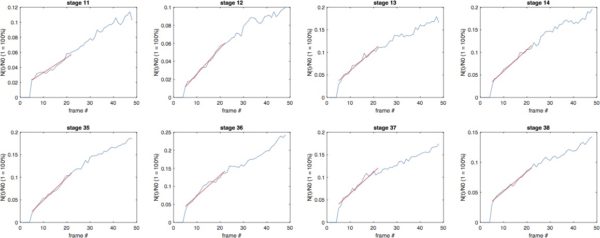
Cell Number Recovery (12 hr):
BdEC/BdEC:
T24/T24:
Wound Healing Stats (based on counting):
Bar: mean. Error bar: standard error. n = 5 x 2 (5 samples, imaged at two non-overlapping FOVs). For T24/T24, n = 4 x 2 (4 samples, imaged at two non-overlapping FOVs).
Wound Healing Stats (based on tracking):
Bar: mean of median values [n specified in top-right-most figure panel]. Error bar: standard error. n = 5 x 2 (5 samples, imaged at two non-overlapping FOVs). For T24/T24, n = 4 x 2 (4 samples, imaged at two non-overlapping FOVs).
It is evident that the BdEC/null and BdEC/BdEC outperformed others in terms of wound healing. T24 wound healing is actually quite impaired in the PECBM+ media. Interesting though, that the T24 is able to proliferate to 100% confluency (from 30% confluency) despite not healing well in the PECBM+ media.
Interesting Factoids: The rate of cellular dispersion (D) can be estimated from the persistent random walk (PRW) model.
where S and P the instantaneous speed and the persistence time predicted by the PRW model. D should be proportional to the wound recovery velocity and the cell-density normalized cell number recovery.
Future Work (To-Do List)
Growth BdEC in PEBCM only, without the corneal epithelial cell growth kit.