Membrane Functionalization: The Pursuit of Chemically Specific Separations
In pursuit of an extremely sensitive biomarker capture sensor, I have been tasked with chemically functionalizing the surface of our membranes. Our end goal is to attach antibodies to the surface of our membranes which can be used to selectively capture target antigens at a wide range of concentrations. Focusing on low concentration ranges in order to make the sensor more physiologically amenable, applications of these sensors are proposed to be within urinary extracellular vesicle capture. A system which could use easily provided urine samples despite their low biomarker concentrations would provide more concrete data about an individuals health. This system could also potentially limit or erase the need of invasive cystoscopy procedures. Overall successful functionalization of our membranes would allow for the exploration of new opportunities for our membranes in both new and old applications.
Epichlorohydrin Functionalization
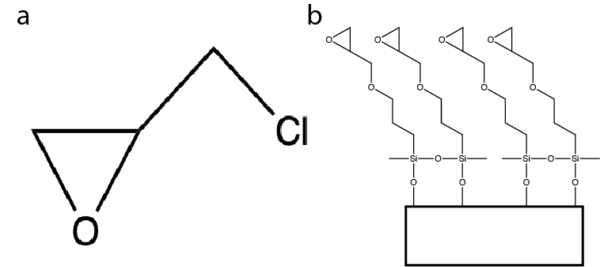
Previously Greg has found that by attaching the molecule epichlorohydrin (chemical structure shown in Figure 1a below) to the surface of our membranes (shown in Figure 1b), we can capture protein G that is tangentially flown through our chips shown here https://trace-bmps.org/tangential-and-flow-through-capture-of-protein-g-on-npn/ . Using this as a starting point, we decided to begin to build our membrane up using this molecule.
As shown in Figure 2 below, the entire process to attach epichlorohydrin to a silicon surface is outlined. This involves oxygen plasma cleaning the chip, hydroxylation of the surface with piranha (a mixture of sulfuric acid and hydrogen peroxide), adding epichlorohydrin as a vapor, and vacuum evacuation to complete the process.

The advantage of binding epichlorohydrin to a silicon surface is the ability to present epoxides available to bind to amines. We planned to use this to our advantage by binding propidium iodide, a fluorophore with two primary amines (shown in Figure 3) to the surface of our functionalized surface.

The goal of binding propidium iodide was to prove that our surfaces were actually functionalized with epichlorohydrin. Unfortunately, fluorescent microscopy showed us that our functionalized membranes did not fluoresce. To determine if this was the fault of epichlorohydrin or if there really was no epichlorohydrin on the surface of our chips, we functionalized more chips but instead attempted to attach BSA and then fluorescently tag the BSA. This also proved unsuccessful, suggesting that we were indeed failing to bind epichlorohydrin to our chips. To ascertain this was the issue we bought new epichlorohydrin, tried different cleaning processes, added ethanolamine as a blocking agent control, and tried a new dye system using alkaline phosphatase and 4-MUP. All of these attempts were unsuccessful ultimately confirming that our issue was with our ability to bind epichlorohydrin to the surface of our chips.
(3-Glycidoxypropyl)Methyldimethoxysilane Functionalization
As an alternative to epichlorohydrin, we looked to (3-glycidoxypropyl)methyldimethoxysilane (GOPMDS). This is a silane modification agent which presents an expoxide and attaches to silicon surfaces similarly to epichlorohydrin. The chemical structure of this molecule is shown in Figure 4 below.

Initially we attempted attaching this molecule and testing it with the three dye systems mentioned above (propidium iodide, BSA and FITC, and alkaline phophatase and 4-MUP). In doing so we found that chips dyed with propidium iodide fluoresced greatly. Unfortunately we also found that chips dyed with BSA/FITC and alkaline phophatase/4-MUP once again did not fluoresce. We interpreted these results as proof that GOPMDS was successfully bound to the chip surfaces, but issues binding larger and more complex molecules were present. With this in mind we were cautious about its ability to bind protein G, but thought there might be a chance that it would work since protein G was smaller and less complex. Because of this we went ahead and attempted its conjugation.
Protein G Conjugation
With GOPMDS functionalized chips we attempted to bind fluorescent protein G to its surface. As shown in Figure 5 below we were actually successful.

In the experiment we compared chips functionalized with GOPMDS and fluorescent protein G, GOPMDS and propidium iodide, GOPMDS, and piranha and plasma. The fluorescence of only the GOPMDS chips bound to fluorescent protein G under FITC excitation proved that protein G was attached to the surface of these chips. Fluorescence of only the propidium iodide dyed chips under Texas Red excitation proved that GOPMDS was successfully bound before protein G was bound to the surface. Chips with only GOPMDS and those that were only piranha and plasma cleaned showed little fluorescence in either excitation range, supporting the conclusions above.
Human IgG Conjugation
After successful binding of protein G to our surface, we moved forward by attempting to attach fluorescent human IgG to the protein G coated surfaces. As shown in Figure 6 below, this proved successful as well.

The bright fluorescence of chips with bound human IgG shown above proved that this fluorescent antibody was present. Compared to chips with similar components with no fluorescent human IgG, it is shown that fluorescence must be imbued by the binding of fluorescent human IgG.
In an attempt to show the importance of both GOPMDS and protein G, we then set up an experiment with chips lacking these components. As shown in Figure 7, instead of our results showing the role of these components we saw that fluorescent human IgG binds to most everything regardless of what is present via non-specific binding. Nonetheless we were able to show potentially interesting data suggesting that protein G allows for ordered binding of antibody to our chips thus increasing the fluorescence of the signal.
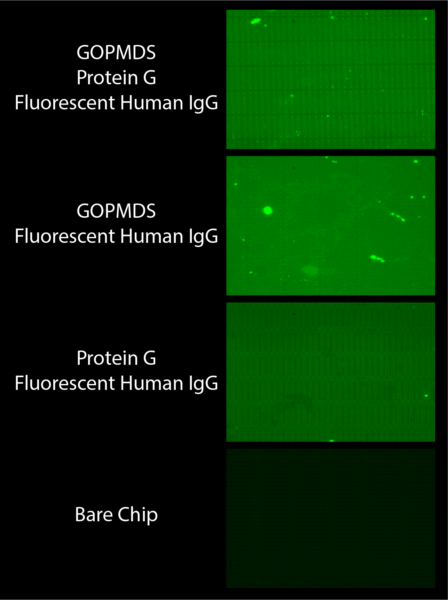
As shown in the picture where GOPMDS and protein G are present, the fluorescence is bright and the slit outlines are sharp. We interpreted this as a result of having ordered bound fluorescent antibody to the surface of the chip due to the presence of protein G. In the chip below this where protein G is absent but GOPMDS is present, similar brightness is shown but the outlines of the slits are lost. We thought this was due to the loss of ordered binding of antibody due to a lack of protein G, but attributed the brightness of signal to the presence of GOPMDS which can still bind antibody in an unordered fashion. The third chip shows less fluorescence and no form around the slits. We thought this was due to nonspecific binding of antibody to the surface of the chip, as protein G would not have bound to the chip in the absence of GOPMDS.
Upcoming Work
With these results in mind, our next step is to continue to try to verify that having protein G coated chips can enable the attachment of antibodies for specific target capture. We are going to do this by working with the Waugh lab and attaching ICAM to the protein G coated chips. Once attached they will verify presence of antibody by rolling cells over these membranes, and I will verify presence of antibody by binding fluorescent anti-ICAM to the antibody coated chips. This will be done in conjunction with a set of chips which are coated in human IgG in order to hopefully show that protein G enables binding of antibody. After this we will attempt to verify antigen capture and test how sensitive this system can be.