The Big Squeeze: Unintentionally Pulling Staphylococcus aureus Through 0.5 µm Microporous Membranes
On our quest to improve the performance of bacterial assays that use the µSiM-CA by modifying our standardized protocol, our most recent modification has caused controversy. Described here, we recently decided to add a step to our protocol where we pull fluorescently tagged Staphylococcus aureus onto the membrane by withdrawing a bacteria filled solution which we load into the well through the membrane. This gave us great success (with regard to bacterial transmission), but concern has lingered that our success may be misleading and our improved performance is actually due to us pulling bacteria through the membrane as opposed to bacterial division. In this post, we set out to test this and apply our newfound knowledge to future protocol iterations.
Initial Pull Through Testing
In our most recent protocol change, we withdrew 50 µL of bacteria filled solution onto the membrane according to the protocol described here. To test whether or not this withdrawal pulls bacteria through the membrane, I replicated this protocol with a few additions to account for the fact that bacteria divides before and after reaching the membrane. To ensure that what we analyzed was in fact bacteria that was pulled through the membrane and not bacteria which merely divided to get there, I set up a waterfall-like system using different sets of µSiM-CAs. This system is described in the protocol below.
Preparing the µSiM-CAs for Testing and Loading them with Bacteria
*Protocol changes shown in red
- Ahead of time, prepare Tryptic Soy Broth (TSB) by adding 30 g of BBL Trypticase Soy Broth to 1 L of purified water
- Mix thoroughly and warm gently until powder is completely dissolved
- Autoclave at 121°C for 15 minutes
- Start an overnight culture by scraping up a chunk of glycerol stock with a pipette tip and adding it to 2 mL of TSB
- Take this overnight culture and put it into a shaker, shaking at 37°C overnight
- The next day, create a subculture by adding 1 mL of your overnight culture to 100 mL of TSB
- Put the subculture in a shaker and let grow at 37°C while shaking until it measures an OD between 0.2-0.4
- While the subculture is growing, thoroughly examine each µSiM-CA device you will use for any defects such as improper sealing between components or broken membranes
- Prepare petri dishes to hold µSiM-CA devices by placing 2 Kim Wipes moistened with sterile dH2O along the perimeter of each petri dish
- Separate and label µSiM-CA devices into two groups: the first used for withdrawing the bacteria onto and potentially through the membrane (group 1) and the second to display any bacteria that is pulled through the first group of devices (group 2)
- Pair the devices so one device from each group corresponds to another in the other group
- Pipette ~15 µL of TSB media by inserting the pipette tip into one of the two open ports of the µSiM-CA and depressing the plunger of the pipette; the media should flow from this port through the bottom channel and out the opposite, open port. Remove the pipette tip before releasing the plunger of the pipette to avoid sucking injected fluid back out of the bottom channel of the device
- Fill the well of the µSiM-CA by pipetting 100 µL of fresh TSB into it; care is taken to not create air bubbles and/or remove them by by withdrawing injected media and injecting it again until no air bubbles are visible
- With the bottom channel and well wet, block one of the open ports using a 3M double-sided tape sticker and stabilize the devices with clamps around its sides
- In the same manner as above, pipette ~40 µL of TSB media into the open port and look to see the well fill with little resistance. If resistance is appreciable, discard the device and prepare a new one
- Remove all liquid from the well (~140 µL), then add 15 µL of TSB into the open port
- Add 35 µL of TSB media to the wells of all of the devices in group 2 so that their total well volume is 50 µL
- Next, add 100 µL of the desired OD subculture to the well of each device in group 1 (the devices used for withdrawal), withdrawing and injecting the subculture again if air bubbles are present until they are gone
- Withdraw 50 µL of fluid from the open port of the devices in group 1, sucking fluid from the well through the membrane, into the bottom channel and out the port
- Immediately take the withdrawn media from the group 1 device and inject this into the well of the corresponding device in group 2
- Replenish the well volume of group 1 devices to 100 µL by pipetting 35 µL of TSB directly into the well
- Allow all devices to incubate at 37°C for 6 hours
After loading, I imaged the group 1 devices at different time points up to 6 hours using confocal imaging and the method described below. Using the same method below, I also imaged the group 2 devices after at least 3 hours of incubating in order to allow any bacteria that may have been pulled through the membrane the chance to divide substantially, increasing our odds of seeing them even if only a handful of bacteria had been pulled through.
Confocal Imaging µSIM-CA Devices Containing GFP Tagged Staphylococcus aureus
- PDMS blocks (~5 mm thick) were laid on top of loaded µSiM-CA devices in order to allow them to be imaged in the microscopy core while maintaining gas permeance
- Any liquid pushed out to the sides of the device is wiped away with a Kim Wipe
- To create an adequate seal, the interface between the PDMS bottom and the acrylic top of the device was sealed with nail polish
- Upon sealing, devices were stored in petri dishes filled with wet Kim Wipes for transport over to river campus
- Once at the microscopy core, images were taken with the 60X water immersion objective and the 1.5X magnifier on the base of the confocal microscope. The 488 nm laser was used with the 525 nm filter
- While imaging, devices were stored in an incubator stage attached the confocal microscope which is held at 37°C
- In between imaging sessions, devices were stored in an oven held at 37°C containing two 1 L beakers of DI water
Results
We tested three devices, using six in total (three for each group). The results are shown in Figure 1 below, indicating that we did in fact pull bacteria through one out of the three devices.
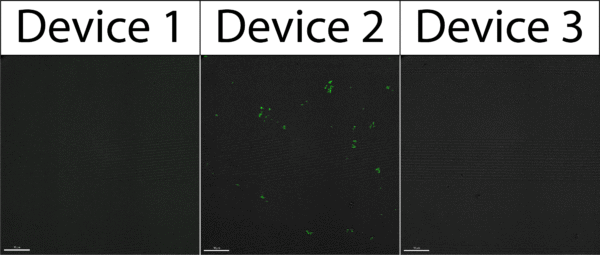
Seeing that we did pull bacteria through the membrane using this protocol, our previous pull down results came into question. What was previously thought unlikely we now had evidence of, with these results suggesting our bacteria has some elasticity to it and a withdrawal of 50 µL provides enough pressure to squeeze the bacteria through our pores. No longer able to trust our previous pull down results, we decided to decrease our pull down volume in hopes of finding the point at which the pressure applied by our withdrawal was small enough where bacteria did not get pulled through the membrane.
Pull Down Titration
Seeing that 50 µL withdrawals caused bacteria to get pulled through the membrane, we decreased our withdrawal volume in steps of 10 µL while following the same protocol described above. Images showing our results are shown in Figure 2 below.
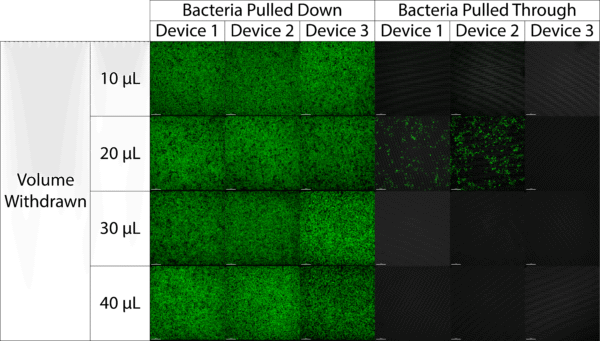
Seen in Figure 2 and shown in Figure 3 below, bacteria was not pulled through using a 10, 30 or 40 µL withdrawals but it was in two devices using a 20 µL withdrawal. Summarized in Figure 3, this means that bacteria was pulled through when using both 20 and 50 µL withdrawals.
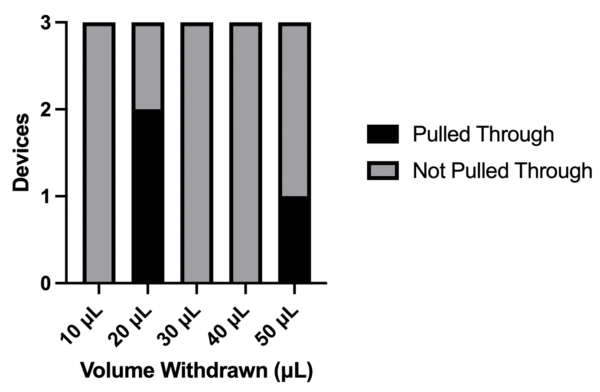
Despite not being pulled through using 30 and 40 µL withdrawals, bacteria getting pulled through at 20 µLs made us believe that pull through events were relatively rare and we must have missed instances where this would occur at 30 and 40 µLs if we had used a higher number of replicates. As a result, we planned to only use 10 µL withdrawals going forward. Acknowledging that the same could be said for 10 µL withdrawals, going forward I planned to continually check to see if we pull bacteria through using this withdrawal until a substantial number of replicates are completed.
Conclusions and Future Directions
Surprisingly, we did find that Staphylococcus aureus is flexible enough and the pressure we applied to the bacteria using our withdrawals was great enough that we were able to pull bacteria through our membranes. This invalidated our previous withdrawal results and led to us titrate our withdrawal volume until we found a volume which did not pull bacteria through our membranes. Finding this volume to be 10 µL, we now plan to attempt to replicate the success we previously had with the withdrawal modification in our future bacterial assays.