HBVP (pericyte) and hCMEC/D3 (endothelial cell) µSiM Coculture Optimization
Introduction
Breakdown of the blood-brain barrier (BBB) is one of the earliest signs of sepsis and is linked with long-term cognitive impairment. We are developing a coculture model of the human blood-brain barrier (µSiM-hBBB) to elucidate mechanisms of BBB breakdown during sepsis. The first version of the µSiM-hBBB will be composed of two cellular components of the neurovascular unit: brain microvascular endothelial cells (BMECs) and pericytes. While BMECs are the main component of the BBB, pericytes are key support cell of the neurovascular unit (NVU), along with astrocytes and microglia. Pericytes not only help stabilize the BBB, but are key responders during inflammation and help guide immune cell transmigration.
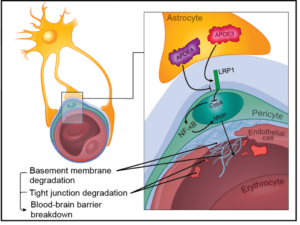
While we will eventually use human induced pluripotent stem cells (hiPSC), we first need to test and optimize coculturing methods on the µSiM. As such, I am testing methods to culture the commercially available hCMEC/D3 endothelial cell line (Millipore) in the top well and human brain vascular pericytes (HBVP, ScienCell) on the top of the bottom channel. This configuration allows the cells to grow 100 nm apart across the nanoporous membrane for good cell communication.
Methods and Results
HBVP (ScienCell) and/or hCMEC/D3 (Millipore) were cultured according to manufacturing protocol seeded in µSiM devices between passages 1-7 and 1-10 respectively. Protocols generally follow ALine Modular Device Cell Culturing: hCMEC/D3and ALine Modular Device Cell Culturing: Bottom Channel Culturing protocols with exceptions described below. Poly-l-lysine (ScienCell) was coated at 2 µg/cm2, collagen type 1 (Sigma) at ~25 µg/cm2, and fibronectin (Corning) at 5 µg/cm2. HBVPs were seeded at 14,000-25,000 cells/cm2(depending on cell availability) and hCMEC/D3 were seeded at 40,000 cells/cm2.
Attempt 1: Coat channel with PLL and well with col.1/fibronectin
Methods
- Coat channel 1 hr with 20 µl poly-l-lysine
- Seed HBVP in pericyte media, flip device (add 80 µl media to top well). Incubate.

- After 2 hours, wash out channel and replace media in well. Flip device to normal orientation (well up) and incubate.
- Next day: coat well 1 hr with collagen type 1/fibronectin mix
- Seed hCMEC/D3 in assay media (AM). Replace pericyte media in channel with AM.
- After 2 hours, wash out well and channel with AM
- Feed cells every 2-3 days with AM, replacing both channel and well
Results
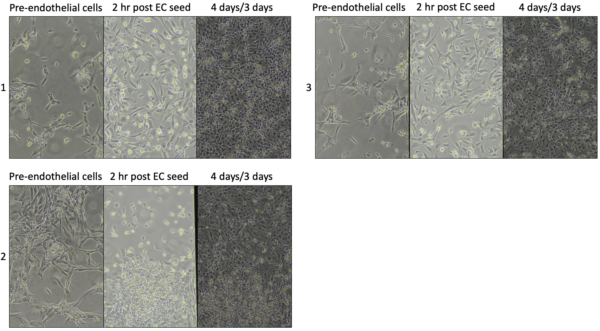
I could easily switch focus between the two cell layers, indicating the pericytes adhered to the bottom of the channel. Also, the hCMEC/D3 looked unhealthy at first. I was surprised they grew to confluency. This may be due to coating the well with collagen/fibronectin after media was already in the well and while pericytes were growing in the channel.
Poly-l-lysine can be almost like glue, and cells may have adhered during the couple minutes before flipping. Next, we will try coating the membrane with not coating with poly-l-lysine.

Attempt 2: No PLL coat
Methods
- Seed HBVP in pericyte media, flip device (add 50 µl media to top well).
- After 2 hours, wash out channel and replace media in well. Flip device to normal orientation (well up).
- Next day: coat well 1 hr with col.1/fn mix (ideally I would have done this before HBVP addition but I did not have time the first day)
- Seed hCMEC/D3 in AM (replace pericyte media in channel with AM)
- After 2 hours, wash out well and channel with AM
- Feed cells every 2-3 days with AM, replacing both channel and well
Results
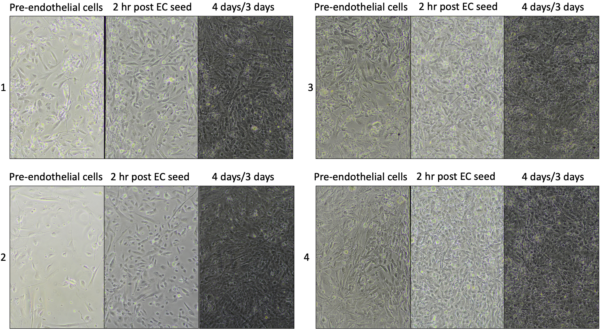
HBVP were able to adhere to the top of the channel and membrane without PLL but the morphology of the cells seems to differ. They did, however, continue to grow over the week of culture and looked healthy when viewing independent of hCMEC/D3 (not pictured). Also, Device 2 cells seemed to have clumped, potentially after pipetting during hCMEC/D3 addition, which could indicate we may need to coat chips for HBVP cultures. Next I will try coating with PLL prior to device assembly.
The cultures in the end were not healthy and made poor monolayers, but this may be due to coating with collagen/fibronectin after media was in the well and while pericytes were growing in the channel. I did not have time to adjust this time, but will coat prior to HBVP addition in the next iteration.
Attempt 3: Coat chip with PLL prior to device assembly
Methods
- Coat trench side of chip 1 hr with 50 µl PLL (will reduce to 20 µl to avoid wicking off chip)

- Remove PLL, wash with water and dry/UV 20 min
- Assemble devices
- Coat well 1 hr with col.1/fn mix. Remove and wash
- Seed HBVP in pericyte media, flip device (add 50 µl media to top well)
- After 2 hours, wash out channel and replace media in well. Flip device to normal orientation (well up)
- Next day: seed hCMEC/D3 in AM (replace pericyte media in channel with AM)
- After 2 hours, wash out well and channel with AM
- Feed cells every 2-3 days AM, replacing both channel and well
Results
This method worked much better, with the pericytes adhering to the top of the channel and no pericytes on the bottom. They stayed on the top of the channel through the 7 days of culturing, without falling to the bottom.
There are also clearly more cells after hCMEC/D3 seeding (more than HBVP growth over 2 hrs, indicating successful attachment of endothelial cells). However, it is hard to distinguish between cells, so in the future we should have HBVP monoculture and hCMEC/D3 monoculture controls to clearly show differences in cell density between culturing conditions. In addition, we may need good imaging or post-hoc analysis to determine coculture health for assays.
Another note, there was variable seeding densities of HBVP and the flipping set up risks media wicking out of the channel when flipped. We will work to improve seeding with low volumes and try clamps to eliminate wicking.
Final note: I did permeability across these cocultures with Dextran-AF488, 10 kDa and saw almost no diffusion across the cell layer over ten minutes (data to come later). We seem to have made a tight barrier.
Attempt 4: Same as attempt 3 but compare coculture to monocultures
Results
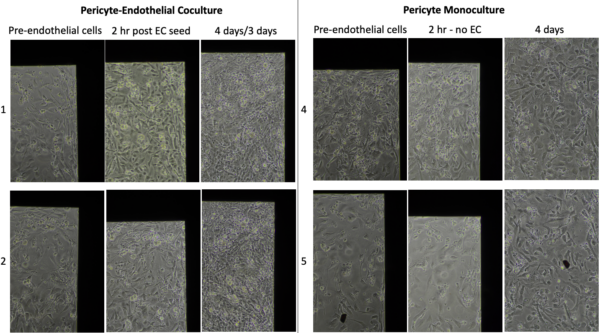
There is clear addition of new cells in coculture with minimal change over 2 hrs in HBVP monoculture. I took images at the top corner to compare the same location over time.

There are clearly more cells in coculture than just from adding hCMEC/D3 to membranes.
I plan to do permeability on these devices to compare the two monolayers and the coculture.
Conclusions
The best coculture protocol for HBVP hCMEC/D3 is to precoat the trench-side of chips with PLL, assemble devices, and then coat the flat side with col.1/fibronectin. Adding pericytes on day 1 and hCMEC/D3 the next day works, although no other methods have been tested. I will continue to plate HBVP and hCMEC alone controls when possible for comparison.
We just received clamps, so I will test those for improvements in flipping the devices for pericyte attachment to the top of the channel. They may also help reduce touching devices with gloves, decreasing the risk of contamination, if we keep devices in the clamps during culturing.
We need to make or buy a chip holders like those pictured for precoating one side of the chip. This method has room for improvement, as sometimes the liquid hits outside of the chip surface and wicks off. This could results in not coating that chip or accidentally coating the opposite side of the chip if the liquid pulls to the underside. This may be a niche issue and can be resolved later. Another potential solution is to try the collagen/fibronectin mix on both sides, and precoat by submerging the cells in that solution prior to assembly. It is unknown if HBVP bind and grow well to this matrix or if the solution would coat the membrane effectively in this fashion. As such, I will only test if necessary.
Update
Immunofluorescence Staining
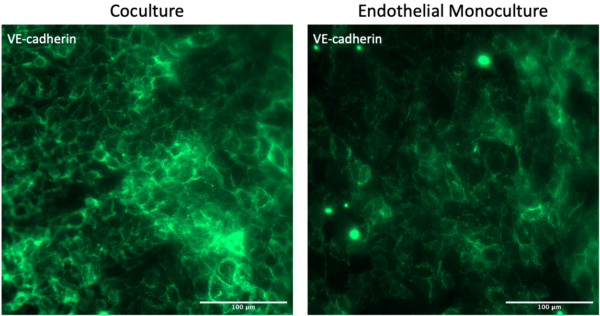

After one week of growth, the VE-cadherin junctions look comparable between coculture and monoculture devices. Obviously we do not see PDGFRb staining in the channel of the monoculture device. Importantly, there is no endothelial cell contamination in the channel from the well.

While there was some background red staining, the pericytes stained strongly for PDGFRb compared to the secondary only background. There is a clear separation of red pericytes and green endothelial cells (see side slices), and there does not appear to be contamination between cellular compartments.
Note that these are representative images but I only had N=2 for coculture and N=1 for monoculture control. We still need to try live cell staining as well.
