Control DNA Experiments
A quick update since we last talked about how DNA controls were proving to be problematic: Things are working again (more or less). I went back top basics, remade most of my solution, ordered new DNA samples, and, most importantly, piranha-cleaned my chips. The first three experiments I did on basic 30-nm thick 5.4mm square chips I made myself, and I got DNA (2000bp, 2000bp, and 5000bp) through each one without too many issues. Whether it was Piranha or something else that made the difference is not clear, but it is a matter of superstition now so I will keep doing it for the time being.
An interesting outcome of these experiment was learning that sample loading technique in the flow cell makes a surprisingly large difference. In our old liquid cells (no flow), I would simply spike in a small amount of concentrated sample and mix it in-situ briefly, and diffusion would take care of mixing. In the flow cell, diffusion happens far too slowly, and sample must be pre-mixed before being injected. Doing so makes an event rate difference of a factor of 10-100, from <0.05Hz event rate to about 1Hz. It’s possible that some of my issues previously were due to this factor.
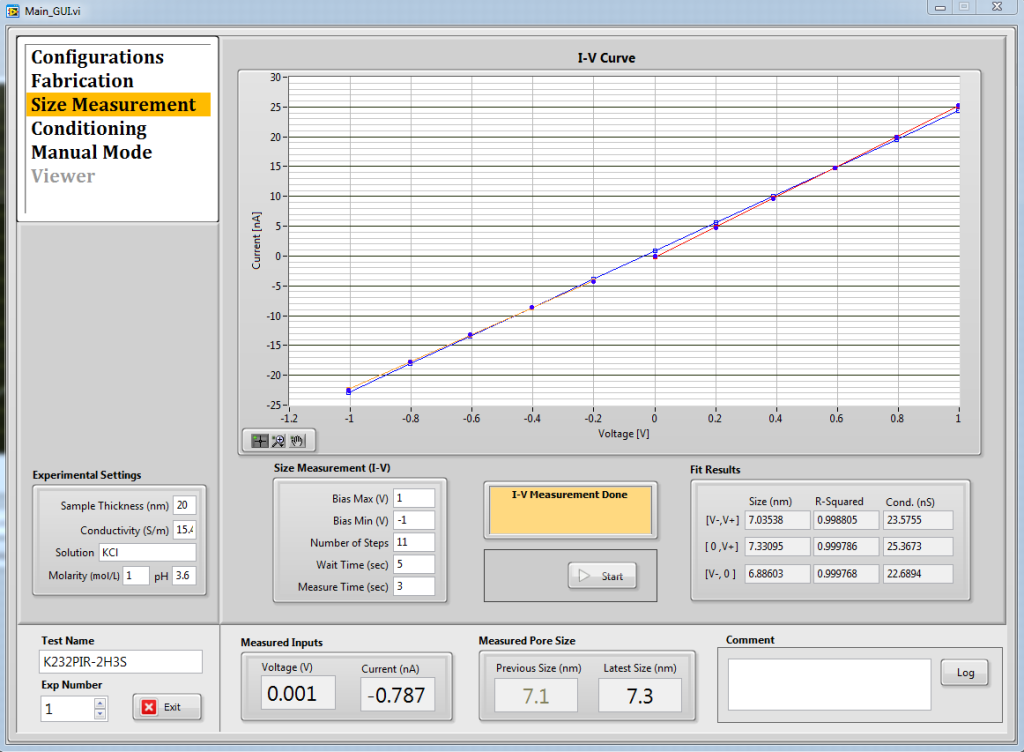
Finally, I ran a control experiment on a chip with the oxide pattern (2-micron holes with 3-micron spacing, no NPN yet) which I Piranha cleaned prior to use, and I was able to see 1000 bp DNA passing the pore. The signals were very odd-looking, but they are clearly DNA and they have a reasonable event rate. Experimental details:
The pore itself was 7-nm in diameter, assuming a 20-nm membrane thickness.
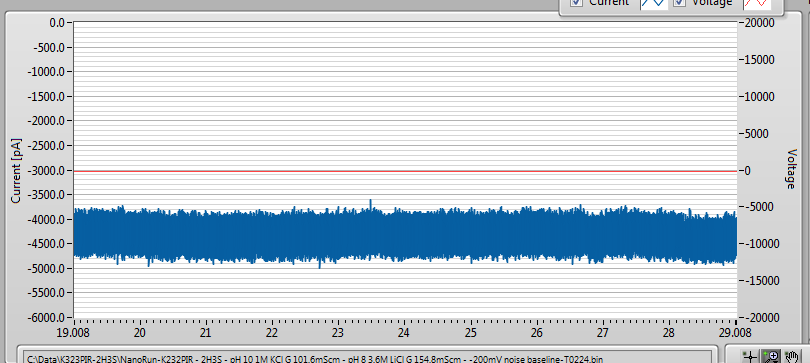
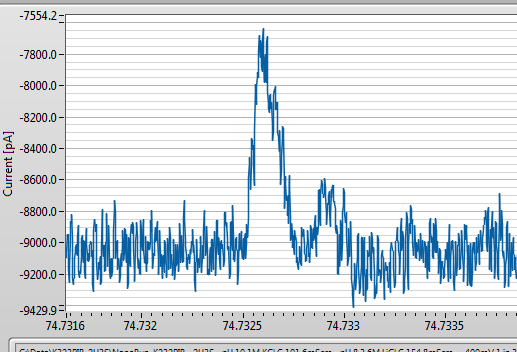
It was a little noisier than I would normally like for a DNA experiment, but since I am running pretty low on these I can’t be too choosy. Here is the baseline (-200mV applied, corresponds to negative bias on the oxide patterned side with the etch pit side grounded). Before adding DNA, we get a relatively high 1/f noise baseline, but no signals that could be mistaken for DNA:
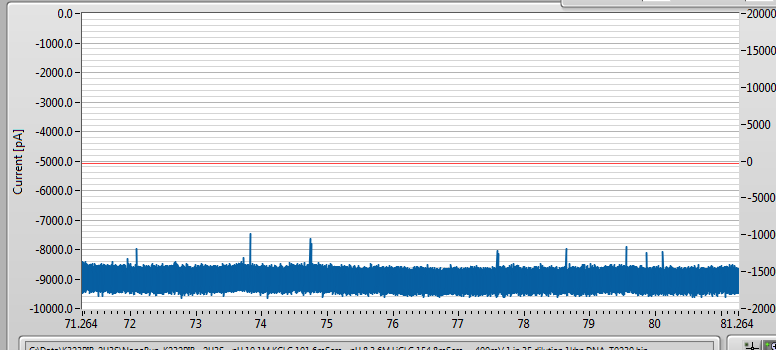
After adding 1000bp DNA at an effective concentration of about 22nM, I get lots of events once I ramped the voltage up to -400mV (-200mV seems to be too low for a good event rate on membranes much thicker than 10-nm, at least so far).
This is where things start to get weird.
I am filtering at 100kHz. Normally, this means there should be a ~10 microsecond rise time for DNA events, where the signal, which should be a square step, gets distorted by the filter. You can read about it in detail here: http://pubs.acs.org/doi/abs/10.1021/nn405761y. Because these DNA molecules are very short, you expect to get a fair amount of attenuation from the filter and some reconstruction is needed to make sense of things, and I’m fine with that since I am really just looking for confirmation that DNA goes through, and I have that. However, the events look like this, close up:
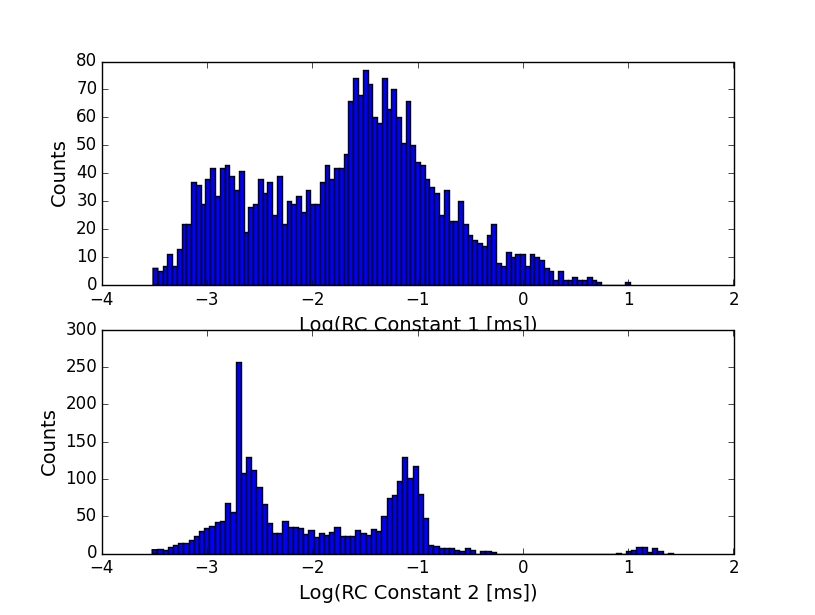
The first thing to notice is that the effective rise time is a lot longer than it should be for this particular filter. In fact, where we expect something on the order of 10 microseconds for the rise time, MOSAIC extracts the following distributions for the rise times:
The peak at ~ -3 is what we expect to see given the filter settings. The higher peak is something I haven’t seen before. The long RC time suggests that something unusual is happening – either the presence of the oxide spacer adds significant access resistance (unlikely, since the 2um holes would have negligible electrical resistance compared to the pore) or for some reason the effective bandwidth is reduced.
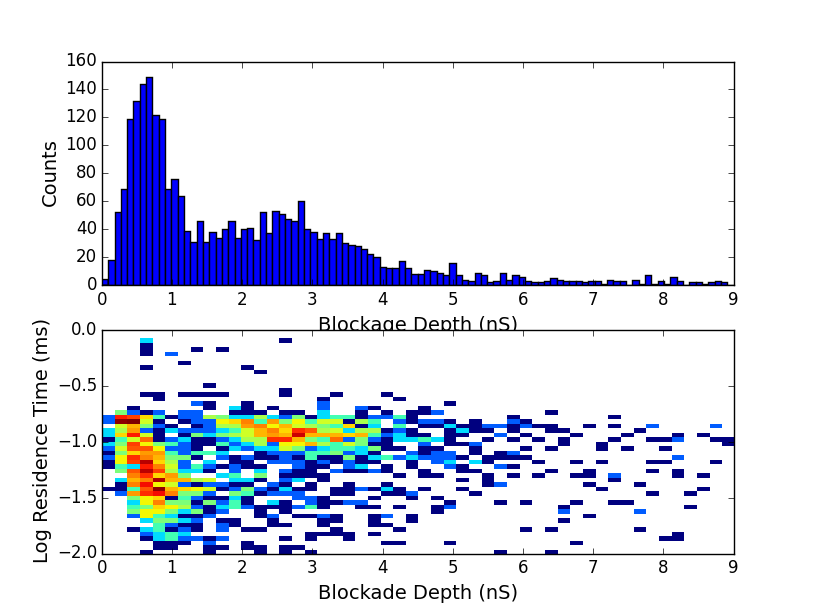
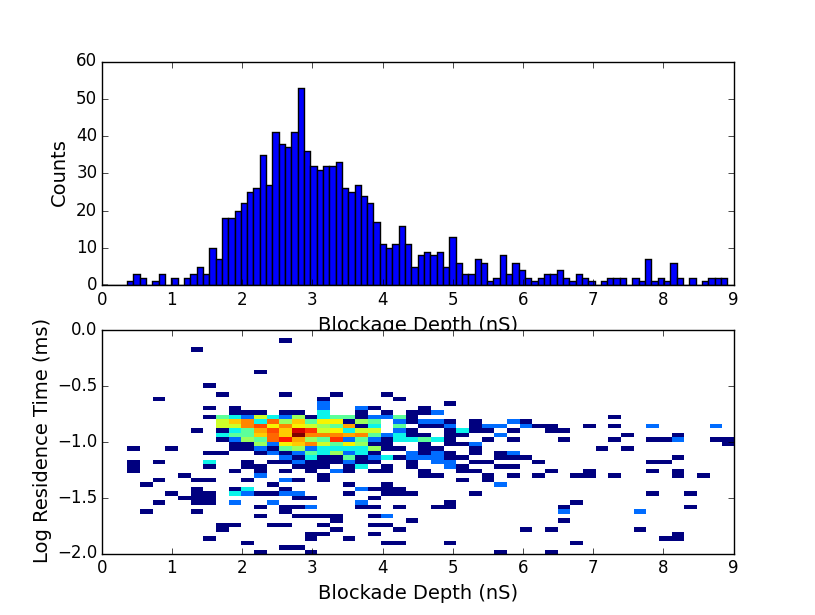
We can look at blockage depths of events when partitioned by RC constant. We expect a blockage of about 3nS for dsDNA in these conditions. Without any partitioning beyond limited RC constants to greater than 0.3 microseconds we get this:
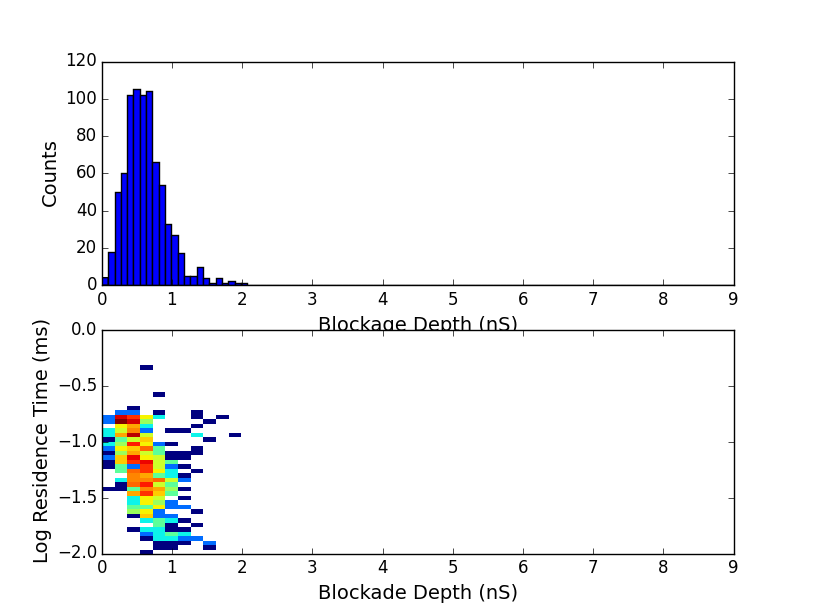
We see a strong peak at 0.5nS and another peak at 3nS as we hoped. Looking only at RC constants between 0.3 and 10 microseconds we get only the first peak:
and looking at RC constants between 10 and 1000 microseconds we get the second peak:
The peak at 0.5 likely corresponds to collisions, where molecules get close enough to the pore to be seen but do not actually enter it. There are usually not nearly so many of those as we see here, suggesting that the oxide spacer might be causing some steric hindrance to DNA passage. But the odd thing is that we can say that molecules which fully translocate the pore have much higher rise times than the filter should cause. My first guess as to why is that the access resistance of the oxide well would be enough to register in the nanopore, since a gradually increasing blockage would potentially mimic an exponential rise time, and the RC constants in question are comparable to the entire translocation time, but some quick calculations show that R_well / R_pore < 0.00005, so this seems unlikely. Another option is that the oxide spacer somehow distorts the shape of the access region to produce this effect (something that could be simulated, if this becomes a trend with these devices). I don’t think it could be that the bandwidth is actually changed, since rise time comparable to the RC time would mean heavily distorted events which would not show the blockage level we are looking for.
However, if that’s not the case, then the usual fitting models might not apply to these oxide devices. I don’t think anyone has explored nanotopography effects on translocation dynamics before, so there might be some interesting work to be done in this area even without the NPN layer.
I am going to run a few more controls (next one planned with 5000bp DNA so that blockages will reach their full height and stay there for a while to facilitate fitting, on a 2um hole/3um space chip). I will also run DNA in the other direction next time to see if the odd behavior happens when DNA moves enters on the opposite side to the pattern. Then I will run a similar experiment on a chip with posts, to see if I can distinguish different nanotopographies from DNA signals. I still have the nanopore device that generated this device mounted, so I will see if I can clean it out and run DNA backwards in that one as well (after a few days, it may be fully clogged).
Experiments that actually have NPN on them will hopefully begin next week, once I iron out my membrane transfer issues, but I think there is enough interesting stuff to explore here that it’s worth spending some time to understand what the oxide layer is doing to translocation dynamics. Or maybe it will turn out to be a quirk of this particular pore. Stay tuned!
Kyle,
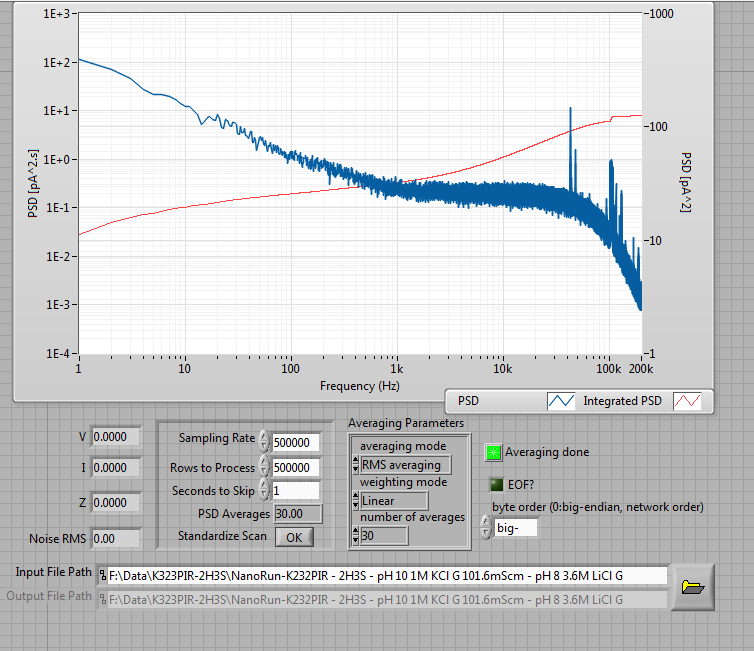
very interesting results. Your PSD is a little surprising. The dielectric/capacitive noise sources do not seem to contribute much at high bandwidth, while there is a relatively high thermal noise floor, which means that R is small… Does your pore size contributes this magnitude of white noise?
I don’t think we can see the true noise floor there – I think the 1/f dominates it past the point where capacitive noise takes over, so we don’t see the flat portion of the PSD clearly.
I’m not sure what inspired you to pre-mix the solutions, but great work solving that issue.
Slower translocation is good, right? Might there be direct physical interactions between DNA and the oxide that are slowing it down? Also, we don’t exactly know where the pore is relative to the oxide. Regardless of the cause (electrical or physical), I suspect the location could change the behavior.
Yes, it’s certainly possible that proximity to the wall of a microwell leads to odd access region dynamics that would lead to effects like this, back of the envelope resistance calculations notwithstanding.
Slow translocations are good, but I should clarify that these events are not really abnormally slow on average. What is really surprising here is the clear presence of two distinct populations of events with a monodisperse sample, as well as the long rise times in the events. This suggests that we can actually sample the presence of the oxide spacer with DNA, something that I would never have anticipated going in, considering the disparity in size scales between the nanopore sensing region and the microwells.
Neat! So the wells themselves potentially are acting as a poor-man’s filter, preventing the quick migration of DNA into the nanopore region.
If the pirranha clean worked effectively, there should be very little irregular native oxide. I wonder if this clean has caused the surface charge to increase (something like silica nanoparticles http://pubs.acs.org/doi/abs/10.1021/la501877v) and is causing the DNA to have more interactions with the walls before entering the nanopore? Or does that even matter because we are using a really high salt concentration which would screen out those effects?
When we are designing chips in the future, we could potentially lower the oxide height, giving the oxide layer less influence, or increase the size of the wells. 2 micron holes is the larger of the two substrates; I wonder if the 1 micron hole substrates will have even more stretched out times.
Alternatively, if the bimodal distribution that you observe is a function of oxide volume, you could try and use the post chip substrates and see if that improves the problem.
I should clarify that when I say long rise time I do not mean long passage time. Rise time refers to the filter response (long rise time means that the apparent effective bandwidth is reduced) but the events are not actually really any longer than I would expect.
I just did another experiment (much less nice, the pore grew the entire time so analysis will be tricky) but it doesn’t look like the long rise times are present in this one, from glancing at a few of them. So either it was a quirk of that pore, or Jim was right and it could be something to do with the nanopore placement relative to the wall. I also tried 5kbp DNA this time and was unable to get any signals at all.
I will try the same thing with the posts and the smaller holes going forward, and see what comes up.
If the oxide height could be reduced or the hole diameter increased while still ensuring no contact between NPN and the membrane that might be ideal going forward.
I would not expect charged walls to play too much of a role due to screening, as you say. Piranha cleaning nanopore chips has been around for a long time, so I think effects like that would have been noted before.