Effect of Exosomes on Wound Healing Assay – 2
Hello Everyone,
So Shannon and i have been working on studying the effect of exosomes in different conditions for wound healing assays and random migrations. This post is about one of our experiments (wound healing) to give you an update on our data and what we’re able to get with Henry’s matlab code.
How the Code Works:
All analysis starts with binary images of the stage which shows only the nuclei stained with SiRDNA. If stage has a wound then a manual outline, or mask, is created by the user that defines the bounds of the wound region. From there, a tracking code is run which shows the path of every nuclei and analyzes for speed, displacement and directionality. Additionally, if it is a stage with a wound, then the code will recognize every cell that leaves the wound mask and contributes to wound healing.
Speed shown is the instantons speed where a speed is calculated for every tracked movement and is averaged over the whole time point.
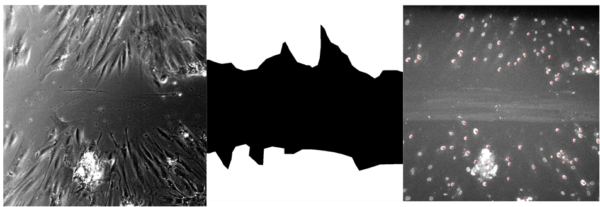

For percent recovery data, the code calculates percent recovery based on cell densities unlike data in most papers, which reports recovery based on area alone. First, the code gets a cell count from the region directly surrounding the wound as a guage or what confluency would be. This count is then divided by the area surrounding to get an initial cell density. The initial density is multiplied by the wound region to get a quantity of cells that would be considered 100% confluent.
The quantity of cells that enter the wound region are tracked at every time point, and this value is divided by the confluent number found above to obtain a ratio of wound recovery. In the case of the bar graph showing recovery percentages for 12 and 24 hours, the ratio of confluency that a stage has reached for that respective time is averaged for each condition.
We designed the experiment with 8 different conditions and the experiment was run for 3 days without changing the media or adding more sirDNA. Here are some side by side videos of different conditions:
- DMEM only [tracking] stage 13
- DMEM + Exosome free FBS [tracking] stage 126
- DMEM + FBS [tracking] stage 118
- ADSC conditioned media (1 Day) [tracking] stage 93
- DMEM + FBS + EGF [tracking] stage 78
- HUVEC conditioned media (1 Day) [tracking] stage 86
- Fibroblast conditioned media (50%) + Fresh media (50%) [tracking] stage 109
- Fibroblast conditioned media (100%) [tracking] stage 103
This experiment was run for 3 days without changing the media or adding more sirDNA. Here are some side by side videos of different conditions:
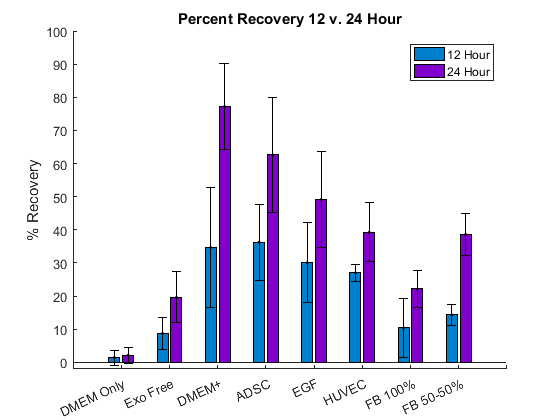
Take home points: (from 12 and 24 hours recovery)
- Cells in DMEM only died immediately, and before getting to 24 hours, cells in Exosome free condition were also dead.
- DMEM + and ADSC are the two best conditions in terms of recovery percentage after 24 hours.
- EGF condition did not show the effect like we expected in terms of 24 hours percentage recovery (Lower than ADSC and DMEM +)
- Fibroblast conditioned media (50-50) showed a better recovery compared to Fibroblasts conditioned media (100%).
- Surprisingly, HUVEC conditioned media showed a good recovery.
Since the experiment was run up to 3 days, we were interested on checking the percentage of recovery over 24 hours.
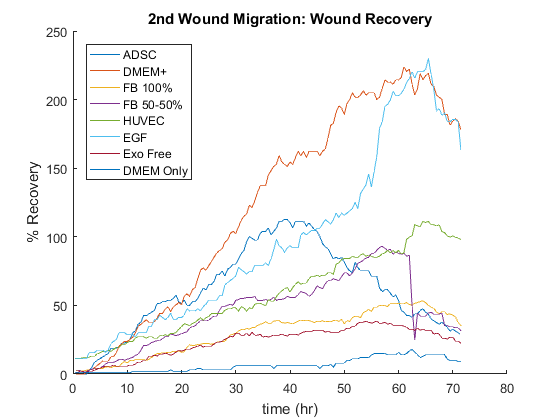
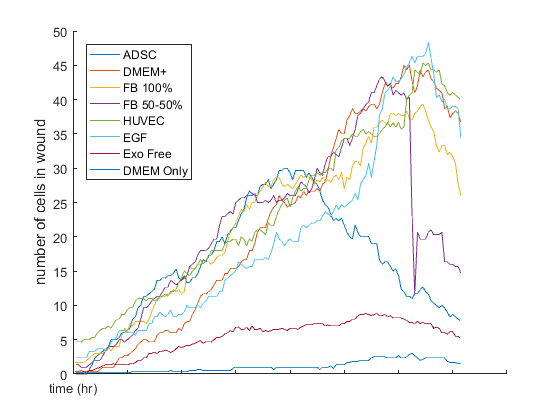
Take home points: (Recovery Plot)
- DMEM + gets to 100% recovery first and ADSC is the second one. It’s important to note that in the first 12 hours, ADSC is leading but at the end of 24 hours, DMEM + is the best condition.
- EGF is not as good as ADSC and DMEM +, but it had a better percentage recovery compared to others.
- Fibroblast conditioned media (50-50) almost got to 100% recovery but Fibroblast 100% is close to 50 percent recovery.
- ADSC drops because it gets super confluent with no room for more cells.
- DMEM + and EGF kept going up on the recovery percentage because they were not super confluent before the wounds were made.
General Take home points:
- The time period that we report our data is important, you can always cherry pick your time period in a way to get your expected results.
- We decided to report the recovery percentage based on the number of cells before wound and after the healing and not based on the area.
- Based on this way of reporting the recovery, it is possible to go over 100 % recovery because of the fact that there is still room for cells to proliferate or even migrate to; specially when the cells before scratching the wounds are not confluent.
- Fibroblast 50 – 50 is better than 100 because fresh media with cow exosomes and nutritions has a good effect on recovery.
- Only DMEM and Exo Free conditions did not survive suggesting that nutritions are important but not as important of exosomes.
- Comparing DMEM + with Exosome free data showed that cow exosomes and possibly other nutritions that were depleted for Exosome free FBS are playing an important rule.
- ADSC, HUVEC and Fibroblasts conditioned media and DMEM + with cow exosomes data show that exosomes are crucial no matter which cell type they are coming from.
Speed Study:
So the next step was to compare the speed of cells migrating to the wound region to the cells from the confluent layers on top and the bottom.
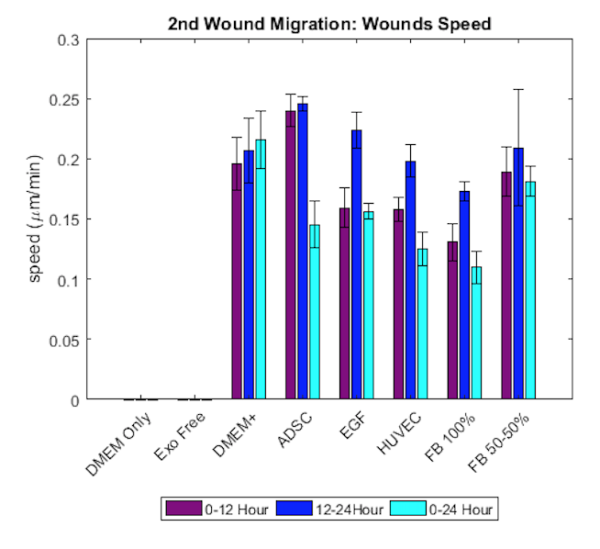
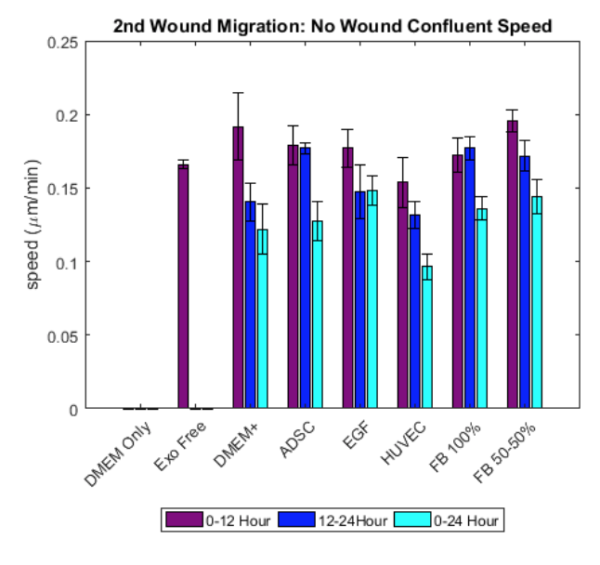
- it seems that speed is not significanytly different between conditions in both the wound region and no wound region.
- Speed in the wound region seems to be higher compared to the speed in the no wound region.
- From the videos, it seems that at the beginning cells are not moving to the wound region and then as soon as they break their bindings to the other cells, they start migrating to the wound region.
- Also as experiment goes on, and the wound is getting healed or closed, the speed drops because more cells are there and there is not enough room for cells to migrate.
- It’s important to pick the time period correctly even for speed for different condition comparisons.
Hi Aslan!
Just a thought but in big-Pharma-type assays it is common to use a well where an object (a strip of silicone film?) is inserted into the well prior to seeding the cells, which are placed in the hemi-wells on both sides. To initiate the “wound” the strip is then removed once the cells are confluent. This greatly aids the morphology and reproducibility of the “scratch” and makes sure that the distance they have to migrate is consistent from run to run. I think a strip of device silicone stick-tioned to the surface would work pretty well..
Kevin-
Thank you so much, and i think i have seen it but i will look it up. We have also made a device that we call wound-maker to make sure that our wounds are similar in different experiments/conditions. You can find the wound maker design in this post: https://trace-bmps.org/wound-maker/
Heh! “wound-maker” sounds like a Viking battle-axe..
Have a look here the kind of thing i was meaning:
https://www.ncbi.nlm.nih.gov/pubmed/29608162
or here:
https://ibidi.com/img/cms/support/AN/AN21_Wound_Healing_Assay.pdf