Utilizing the μSIM to model brain endothelial barrier changes in Cerebral Amyloid Angiopathy
Introduction
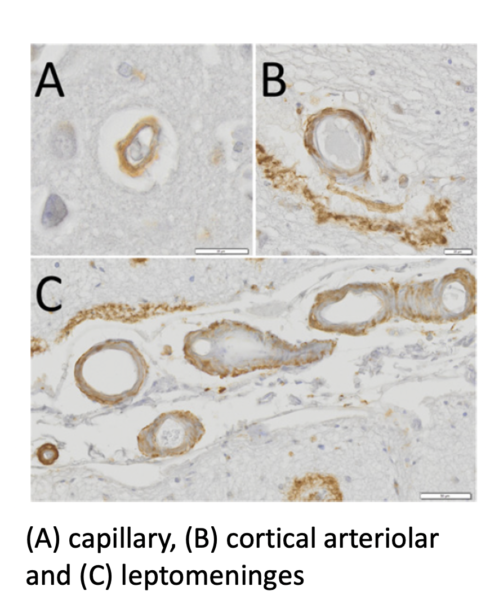
Vascular hyperpermeability and hemorrhagic stroke are hallmark symptoms of Cerebral Amyloid Angiopathy (CAA), a cerebrovascular disease characterized by amyloid-β deposition around the brain microvasculature. In CAA pathogenesis, vascular Aβ accumulation contributes to structural alterations and loss of integrity in the neurovascular unit/blood-brain barrier (NVU/BBB) resulting in “leakier” blood vessels, intracerebral hemorrhages, increased risk of stroke, and cognitive impairment. Vascular Aβ deposition is known to promote junctional protein changes in the brain endothelium fueling the hyperpermeability, perivascular response, capillary deterioration and often occlusion, and blood vessel ruptures (microhemorrhages). However, it remains true that the field lacks a better in vitro model to elucidate underlying molecular mechanisms triggered by Amyloid-β (Aβ) which contribute to the BBB injury and hyperpermeability in CAA. To fill this gap, we are using the μSIM barrier platform to recapitulate amyloid-β interaction with microvascular endothelial cells to further investigate junctional changes. Specifically, my project aims to elucidate how altered expression of junctional protein connexin-43 (Cx43) modified function of Cx43 structures, gap junction and hemichannels and drive the BBB injury/hyperpermeability in CAA disease pathology.
Methods
In this protocol, we are using C57BL/6 Mouse Primary Brain Microvascular Endothelial Cells purchased from Cell Biologics.
Preparing Cells for plating on Device
- Coat top chamber/well with 100 μl of gelatin coating solution. Incubate devices in incubator for 30 mins at 37ºC
- Remove gelatin from top chamber and add complete endothelial cell medium to bottom well
- Seed 120,000-150,000 cells/mL per devices depending on how fast you want cells to reach confluency
- Return devices to incubator after plating cells and make sure to check for cell attachment next day.
- Change medium in both chambers every other day
- A mature barrier to form between cells takes 3-5 days depending on the seeding density
Amyloid-β Treatment
- Cells should be around ~100 % confluent when you begin treatment (Cells need at least a minimum of 3 days after plating to establish barrier)
- Amyloid-β 1-40 and 1-42 HFIP-treated peptides (obtained from Bachem) are prepared in advance using a previously published method (Rosensweig et al, Methods Mol Biol., 2012). Protein Concentration of aggregated peptides (Requires 3 days or ~72 hours of incubation at 37ºC shaking at ~260 rpm) can be determined running a BCA assay
- Amyloid-β 1-40 or 1-42 are diluted to 5 μM concentrations using serum-free medium and applied to bottom chamber
- Top well should be replaced with 100 μl of serum-free medium
- Incubate at 37ºC for 6 or 24 hours
- Follow-up with immunofluorescence protocol for protein of interest
Live Cell Imaging in μSIM :
Hemichannel Functional Assay
Materials to prepare
- HBSS Solution
- Mix HBSS w/ no phenol red with 1M HEPES to final concentration of 20 mM (ex: 1 mL 1M HEPES for 50 mL final volume)
- HBSS-Ca2+ free solution
- Mix HBSS devoid of calcium and magnesium with 1M HEPES to final concentration of 20 mM, MgCl2 to final concentration of 1mM, and EGTA to final concentration of 0.1 mM
- HBSS-Ca2+ free stock with 10 mM Lucifer Yellow (Final Concentration used: 2 mM)
- Prepare cells to be ~100 % confluent when you begin treatment (Cells need at least a minimum of 3 days after plating to establish barrier)
- Cells should be washed twice with warmed HBSS solution, then once with warm HBSS-Ca2+ free solution
- For negative control, only HBSS should be used
- After last wash, ensure that you removed all calcium containing HBSS, if needed, wash an additional time with the HBSS-Ca2+ free solution
- When ready, remove the HBSS-Ca2+ free solution and loaded cells with pre-warmed HBSS-Ca2+ free stock with Lucifer Yellow for 10 mins at RT
- For negative controls, you should prepare pre-warmed HBSS with calcium and lucifer yellow
- Remove HBSS-Ca2+ free with Lucifer Yellow solution
- Then wash cells twice with warm HBSS solution with calcium to induce hemichannel closure
- Images should be acquired immediately after washing steps
NOTES
- During the experiments, you need to handle cells carefully, as hemichannels are mechanosensitive. Excessive shaking and vigorous exchange of media will induce hemichannel opening and therefore increase the background for subsequent measurements.
- For analysis of lucifer yellow uptake, we will either be calculating number of LY-positive cells or measuring total fluorescence intensity
Fluorescence Recovery After Photobleaching Protocol
We used the following protocol: https://experiments.springernature.com/articles/10.1007/978-1-4939-3664-9_12
Results
Utilization of the μSIM device in our experiments provides a tool to study abluminal treatment of amyloid-β on endothelial cells that allows us to obtain high-resolution imaging of junctional proteins that is not possible with traditional transwell.

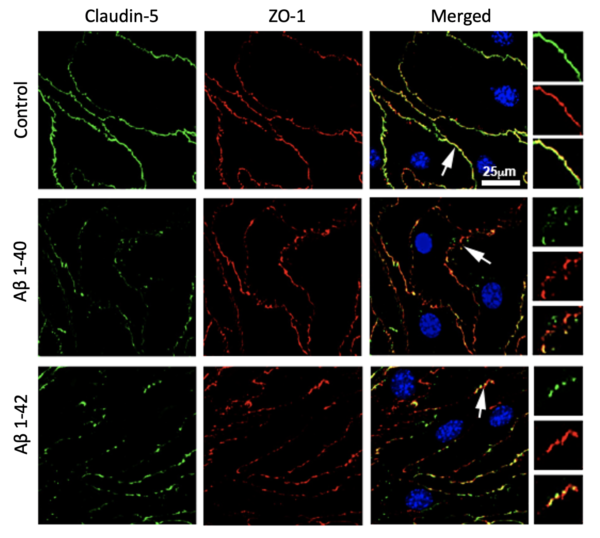
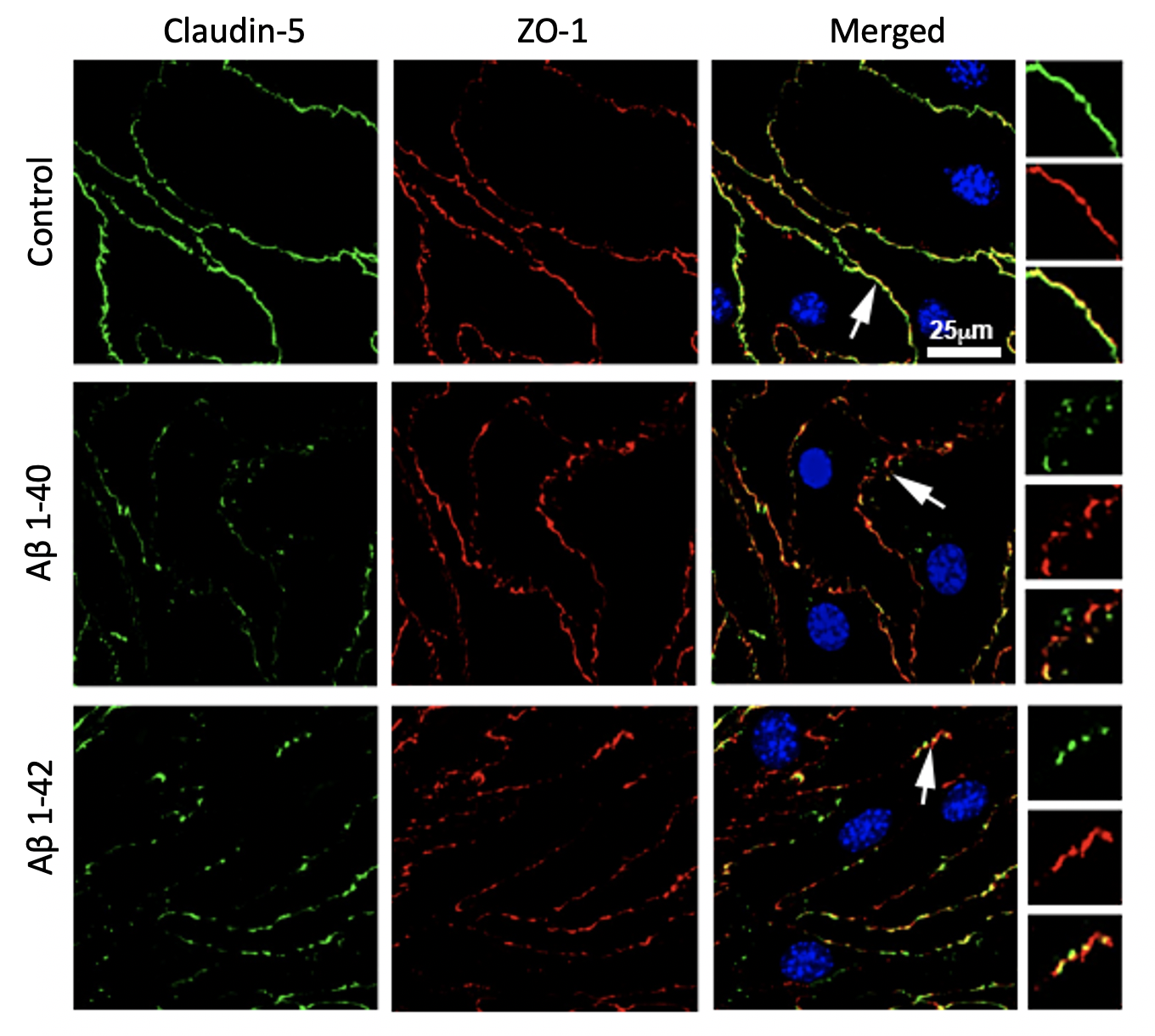
Our immunofluorescence staining in Figure 2 demonstrates that cells exposed to amyloid-β peptides in the lower chamber (i.e., abluminal side) has more fragment staining of ZO-1 and Claudin-5, two junctional proteins essential for establishing a stable barrier, relative to the control cells. We also have previous data from functional assay done in the conventional transwell that showed increased permeability. All this indicates that the tight junction complex is less stable in amyloid-β treated cells. We are also in the process of analyzing junctional protein morphology using an open-sourced Junctional Analyzer Program (Gray et al., 2019).
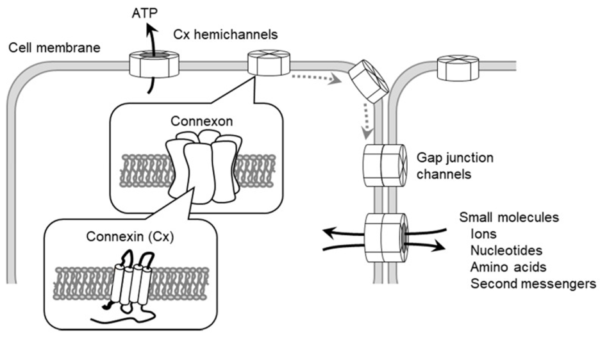
Emerging evidence has demonstrated that modifications in expression and function of other junction proteins like connexins (Cx) can drive changes in TJ complex stability and consequently regulate BBB integrity. Connexins are transmembrane proteins that form a hexamer called a connexon with canonically functions as either a gap junction (GJ) channel allowing intercellular communication or hemichannels (HCs) localized to the cell surface, allowing extracellular communication (Shown in Figure 3). One of the most abundant connexins in the brain is Cx43, which we are interested in elucidating its role in barrier integrity within BECs exposed to amyloid-β plaques. Our staining of Cx43 and ZO-1 suggests an increased expression of Cx43 plaques in amyloid-β treated cells that is then no longer presented when treated with mimetic peptides, Gap 19 and Gap 27, which block hemichannels and gap junctions, respectively.

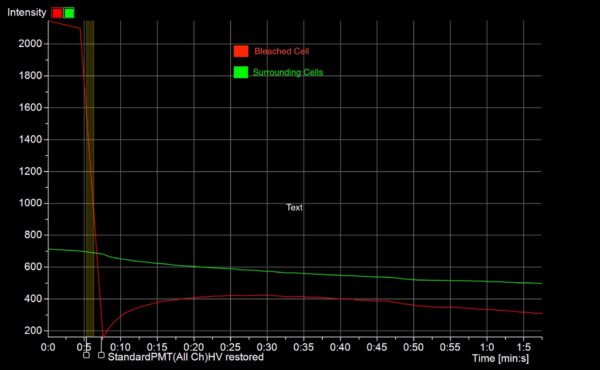
Live Cell Imaging Results

Future Plans
- Duo-Link assay to examine Cldn-5 and ZO-1 interaction under amyloid-β exposure
- STORM Imaging to analyze Cldn-5 strands
- In process of preparing for pericyte co-culture with endothelial cells to further study Cx43 role in CAA-associated injury of brain endothelial cells

Wonderful contribution Muyu. Thank you! Please upload higher resolution versions of the images after today’s meeting. We do this by take screen shots of zoomed in versions and resizing in the post with the editor. I’ll save my science questions for the meeting!