Measuring Protein Contamination of Exosomes Using XPS
One of the major goals of my work with capturing exosomes is to show that our purified population is less contaminated with protein than exosomes purified by ultracentrifugation. This is important because the protein (and all the other junk floating in plasma such as cfDNA) can cause problems with identification and characterization of the exosomes. While there are many protein assays to characterize the amounts of protein in a sample, Jim suggested (and I liked because I’ve had experience with it) using x-ray photoelectron spectroscopy (XPS) to quantify the thickness of a protein layer on the surface of our membranes. Like Auger, XPS is a surface analysis technique that returns signal from the first 10 nm or so of the surface.
To start this study, I basically wanted to show that the amount of protein in a sample with normal flow (also analogous to two pump flow) was higher than a simple, single-pump tangential flow system. As a control, I ran a clean membrane. The spectra for the clean membrane is shown in Figure 1.
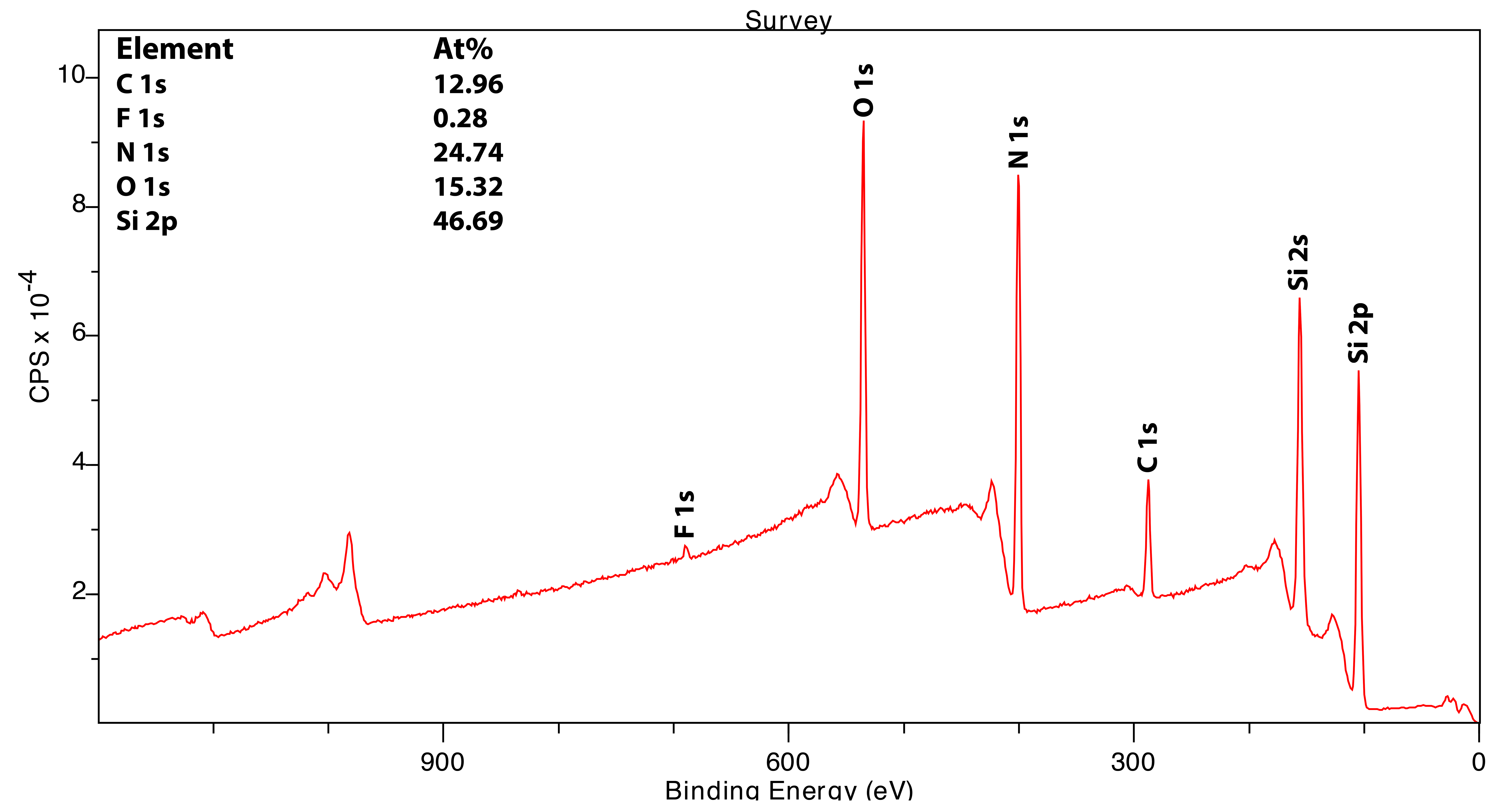
Figure 1: XPS spectra of a clean membrane (from wafer 1255). To note are the fluorine peak (likely from the etching process) and the very strong silicon peaks (both the 2s and 2p peaks are visible).
With a reference for the membrane, I then ran a membrane that has been spun in a SepCon column with plasma and a membrane that was exposed to single-pump tangential flow. One cool thing that I think I should point out is that I was able to run a single slot membrane in the SepCon system without the membrane breaking (2500 rpm for 10 minutes). The spectra for the plasma samples is shown in Figure 2.
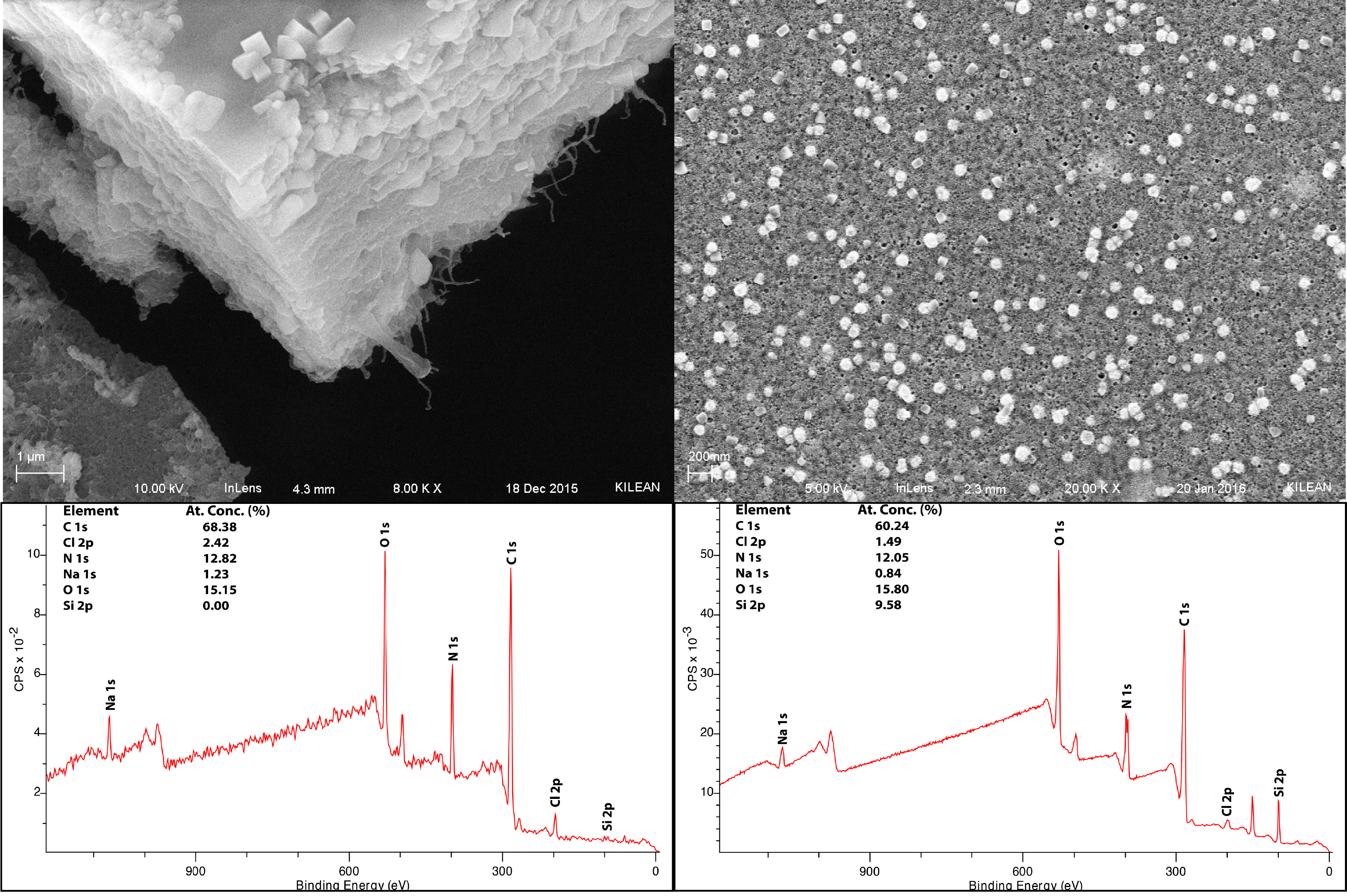
Figure 2: Comparison of protein content for normal flow (left) and tangential flow (right) plasma samples. Note how in the normal flow sample there is no silicon peak and in the tangential flow sample the silicon 2p peak is visible. This indicates that the protein layer is greater than 10 nm in the normal flow sample, while it is less than 10 nm in the tangential flow sample.
What we learned form this experiment is that we can differentiate between normal and tangential flow samples using XPS only. In the future, I will be doing angle resolved XPS to determine the actual thickness of the protein layer (similar to the work done by Jim and Alex Shestopolov in their recent paper). I will also compare the two pump system and quantify the amount of protein in an ultracentrifuged sample. I was planning on having that data in this post, but the XPS system is down unfortunately and I am working on bringing it back online.
On Imgur, they have a cat tax or something at the end of a post, but today I’ll leave you with an exosome tax from one of my recent runs with the purified exosome samples that I purchased:
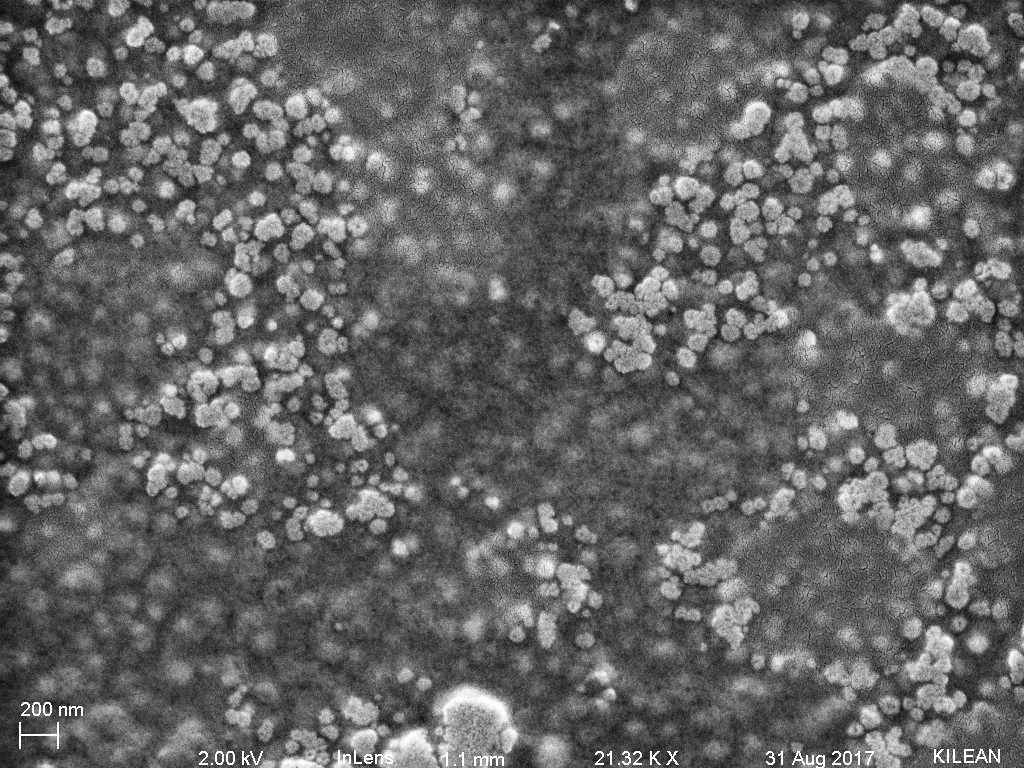

Figure 3: Purified exosomes captured on a membrane (wafer 1228) in tangential flow from a purified exosome stock (HansaBioMed) of 1 x 10^10 exosomes/mL. The exosomes are at a very high concentration and thus stacked on each other and also covered by a thin layer of protein (?) which burned when the electron beam resided on the area for too long.